긴꼬리투구새우 (Triops longicaudatus)
|
|
▣ 국 명: 긴꼬리투구새우 ▣ 영 명: longtail tadpole shrimp ▣ 학 명: Triops longicaudatus ▣ 지정번호: 멸종위기해제종 ▣ 계 통: 갑각강-배각목-긴꼬리투구새우과 |
◎ 생김새 : 꼬리부분을 포함한 전체 길이는 3~5cm정도이며 꼬리를 제외한 몸길이는 2~3cm이다. 등쪽에 몸의 2/3를 덮은 납작한 투구 모양의 갑각을 갖고 있으며 황색을 띤 갈색 또는 갈색을 띈다. 갑각의 앞쪽 중앙에 1쌍의 눈이 있다. 37개 정도의 마디로 나누어져 있으며 가슴에 11쌍, 배에 18~19쌍의 다리가 있고 꼬리 마디 끝에 1쌍의 긴 부속지를 갖는다.
◎ 서식지 : 얕은 민물의 웅덩이나 논에서 서식한다.
◎ 먹이 : 잡식성으로 조류, 유기물, 원생동물 또는 모기 유충이나 자기보다 작은 곤충 등도 먹을 뿐 아니라 포식성이 강해 동족끼리도 잡아먹기도 한다.
◎ 번식 : 주로 단위 생식을 하며 드물게 유성생식을 하기도 한다. 대부분의 개체군내 성비가 편향되어 있다. 다리가 변형된 알주머니에 검붉은 색의 알을 품고 있다가 모래나 흙을 파서 알을 낳는다. 단단한 껍질에 쌓인 알 상태로 가뭄이나 겨울을 나고 환경이 좋아지면 부화하는 특징을 보인다. 유기물이 부족한 물, 햇빛, 22~30℃의 온도 조건에서 부화한다.
◎ 분포 : 아메리카 대륙 및 서인도 제도, 아시아에 걸쳐 널리 분포하고 있으며 한국의 경우 전국의 벼농사지역에서 널리 발견되고 있다.
1) 채집정보
- 채집허가: (멸종위기해제종)
- 채 집 자: 경북대학교 황의욱
2) 채집현황 및 샘플처리
|
긴꼬리투구새우 채집현황 |
샘플처리 |
|||
|
채집일 |
개체 수 |
DNAseq |
RNAseq |
표본 |
|
2014.06.10 |
42 EA |
30 EA |
10 EA |
2 |
3) QC Data
|
샘플종 |
DNA QC |
RNA QC |
Result |
||
|
Concentration (ug/ul) |
Total Conc. (ug) |
Concentration (ug/ul) |
RIN (>7) |
||
|
긴꼬리투구새우 |
81.2 |
6.5 |
757.1 |
9.7 |
pass |
4) 기존 NCBI Taxonomy 유전정보 현황
|
유전정보 종류 |
Triops 속 유전자원 |
Triops longicaudatus 유전자원 |
|
Nucleotide |
5,834 |
405 |
|
Protein |
1,302 |
427 |
|
Genome |
4 |
1 |
|
Gene |
100 |
13 |
5) Sequencing 분석 결과
- Illumina DNA Sequncing 결과
|
Input information |
|||||
|
|
|
|
Region1 |
Region2 |
Total |
|
raw data |
|||||
|
Number Of Reads |
|
|
142,992,046 |
142,992,046 |
285,984,092 |
|
Number Of Bases |
|
|
18,016,997,796 |
18,016,997,796 |
36,033,995,592 |
|
error correction |
|||||
|
Number Of Reads |
|
|
133,578,956 |
131,662,333 |
265,241,289 |
|
Number Of Bases |
|
|
16,566,233,178 |
16,215,232,816 |
32,781,465,994 |
|
pair reads |
|||||
|
Number Of Reads |
|
|
|
|
260,946,666 |
|
Number Of Bases |
|
|
|
|
32,325,997,283 |
|
single reads |
|||||
|
Number Of Reads |
|
|
|
|
4,294,623 |
|
Number Of Bases |
|
|
|
|
455,468,711 |
|
Assembly Results |
|||||
|
Scaffold Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Scaffolds |
212,039 |
212,039 |
52,752 |
28,482 |
14,317 |
|
Number Of Bases |
175,883,178 |
175,883,178 |
144,634,329 |
127,671,924 |
107,976,378 |
|
Avg. Scaffold Size |
829 |
829 |
2,741 |
4,482 |
7,541 |
|
N50 Scaffold Size |
4,075 |
4,075 |
6,997 |
9,227 |
12,212 |
|
N80 Scaffold Size |
594 |
594 |
1,528 |
2,451 |
4,278 |
|
N90 Scaffold Size |
251 |
251 |
906 |
1,571 |
2,933 |
|
largest Scaffold Size |
197,609 |
197,609 |
197,609 |
197,609 |
197,609 |
|
Contig Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Contigs |
1,820,833 |
414,399 |
49,067 |
23,449 |
13,344 |
|
Number Of Bases |
247,327,299 |
178,504,593 |
112,067,821 |
94,666,770 |
80,711,822 |
|
Avg. Contig Size |
135 |
430 |
2,283 |
4,037 |
6,048 |
|
N50 Contig Size |
342 |
1,299 |
5,117 |
6,400 |
7,666 |
|
N80 Contig Size |
63 |
203 |
1,276 |
2,541 |
3,921 |
|
N90 Contig Size |
42 |
137 |
760 |
1,594 |
2,916 |
|
Largest Contig Size |
80,995 |
80,995 |
80,995 |
80,995 |
80,995 |
- Illumina RNA Sequncing 결과
|
Input information |
|||||
|
|
|
|
Region1 |
Region2 |
Total |
|
raw data |
|||||
|
Number Of Reads |
|
|
161,659,804 |
161,659,804 |
323,319,608 |
|
Number Of Bases |
|
|
20,369,135,304 |
20,369,135,304 |
40,738,270,608 |
|
error correction |
|||||
|
Number Of Reads |
|
|
159,867,931 |
159,142,254 |
319,010,185 |
|
Number Of Bases |
|
|
19,943,723,063 |
19,834,516,297 |
39,778,239,360 |
|
pair reads |
|||||
|
Number Of Reads |
|
|
|
|
317,586,644 |
|
Number Of Bases |
|
|
|
|
39,619,172,037 |
|
single reads |
|||||
|
Number Of Reads |
|
|
|
|
1,423,541 |
|
Number Of Bases |
|
|
|
|
159,067,323 |
|
Assembly Results |
|||||
|
Scaffold Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Scaffolds |
128,900 |
128,900 |
22,757 |
8,984 |
2,747 |
|
Number Of Bases |
48,943,255 |
48,943,255 |
27,253,404 |
17,765,153 |
9,245,545 |
|
Avg. Scaffold Size |
379 |
379 |
1,197 |
1,977 |
3,365 |
|
N50 Scaffold Size |
607 |
607 |
1,370 |
2,070 |
3,298 |
|
N80 Scaffold Size |
213 |
213 |
738 |
1,311 |
2,413 |
|
N90 Scaffold Size |
146 |
146 |
605 |
1,143 |
2,193 |
|
largest Scaffold Size |
21,871 |
21,871 |
21,871 |
21,871 |
21,871 |
|
Contig Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Contigs |
1,551,396 |
167,786 |
20,645 |
6,481 |
1,547 |
|
Number Of Bases |
113,674,340 |
49,638,294 |
21,044,559 |
11,374,123 |
4,735,833 |
|
Avg. Contig Size |
73 |
295 |
1,019 |
1,754 |
3,061 |
|
N50 Contig Size |
63 |
394 |
1,074 |
1,762 |
2,936 |
|
N80 Contig Size |
39 |
167 |
662 |
1,222 |
2,269 |
|
N90 Contig Size |
32 |
128 |
574 |
1,105 |
2,124 |
|
Largest Contig Size |
15,032 |
15,032 |
15,032 |
15,032 |
15,032 |
(1) 고품질 genomic DNA 채취 및 표본확보
|
검수용 샘플 DNA 농도 및 총 양, 확증표본 현황표 |
|||||||
|
Sample명 |
농도(ng/ul) |
Volume(ul) |
DNA 총량(ug) |
O.D A260/280 |
확증표본 |
채집허가일 |
채집일 |
|
긴꼬리투구새우 |
285 |
996 |
283 |
1.84 |
○ |
고유종 |
2014.6.10 |
(2) 유전체 서열확보 : 현재 20종 확보
|
Sample |
구분 |
raw data size |
trim data size |
비고 |
|
긴꼬리투구새우 |
DNA |
36,033,995,592 |
32,781,465,994 |
|
|
RNA |
40,738,270,608 |
39,778,239,360 |
|
(3) 기타분석
A. Genome Size estimation
|
샘 플 명 |
Kmer size |
Kmer total |
Pk depth |
Genome_size |
Coverage(X) |
|
긴꼬리투구새우 |
23 |
22,306,759,176 |
79 |
282,364,040 |
101.282 |
B. 반복서열
- 시퀀싱이 완료된 긴꼬리투구새우 DNA 및 RNA의 contigs 중 1kb 이상의 contigs의 반복서열을 탐색(Left; Repeat masking results of DNA contigs, Right; Repeat masking results of RNA contigs).
|
sequences: 23,449 |
|
sequences: 6,481 |
||||||||
|
total length: 94,666,770 bp (94,666,770 bp excl N/X-runs) |
|
total length: 11,374,123 bp (11,374,123 bp excl N/X-runs) |
||||||||
|
GC level: 41.93% |
|
GC level: 46.82% |
||||||||
|
bases masked: 23,664 bp (0.02%) |
|
bases masked: 1,929 bp (0.02%) |
||||||||
|
|
number of elements |
length occupied |
percentage of sequence |
|
|
number of elements |
length occupied |
percentage of sequence |
||
|
SINEs: |
13 |
509 bp |
0.00% |
|
SINEs: |
1 |
47 bp |
0.00% |
||
|
|
ALUs |
0 |
0 bp |
0.00% |
|
|
ALUs |
0 |
0 bp |
0.00% |
|
|
MIRs |
2 |
105 bp |
0.00% |
|
|
MIRs |
0 |
0 bp |
0.00% |
|
LINEs: |
56 |
3,128 bp |
0.00% |
|
LINEs: |
11 |
760 bp |
0.01% |
||
|
|
LINE1 |
13 |
707 bp |
0.00% |
|
|
LINE1 |
0 |
0 bp |
0.00% |
|
|
LINE2 |
9 |
434 bp |
0.00% |
|
|
LINE2 |
2 |
88 bp |
0.00% |
|
|
L3/CR1 |
22 |
1,106 bp |
0.00% |
|
|
L3/CR1 |
5 |
320 bp |
0.00% |
|
LTR elements: |
14 |
1,134 bp |
0.00% |
|
LTR elements: |
3 |
156 bp |
0.00% |
||
|
|
ERVL |
3 |
143 bp |
0.00% |
|
|
ERVL |
2 |
97 bp |
0.00% |
|
|
ERVL-MaLRs |
0 |
0 bp |
0.00% |
|
|
ERVL-MaLRs |
0 |
0 bp |
0.00% |
|
|
ERV_classI |
6 |
309 bp |
0.00% |
|
|
ERV_classI |
0 |
0 bp |
0.00% |
|
|
ERV_classII |
0 |
0 bp |
0.00% |
|
|
ERV_classII |
0 |
0 bp |
0.00% |
|
DNA elements: |
72 |
7,059 bp |
0.01% |
|
DNA elements: |
8 |
775 bp |
0.01% |
||
|
|
hAT-Charlie |
28 |
3,736 bp |
0.00% |
|
|
hAT-Charlie |
1 |
150 bp |
0.00% |
|
|
TcMar-Tigger |
17 |
1,606 bp |
0.00% |
|
|
TcMar-Tigger |
3 |
371 bp |
0.00% |
|
Unclassified: |
2 |
133 bp |
0.00% |
|
Unclassified: |
1 |
72 bp |
0.00% |
||
|
Total interspersed repeats: |
|
11,963 bp |
0.01% |
|
Total interspersed repeats: |
|
1,810 bp |
0.02% |
||
|
Small RNA: |
184 |
11,735 bp |
0.01% |
|
Small RNA: |
3 |
166 bp |
0.00% |
||
|
Satellites: |
1 |
47 bp |
0.00% |
|
Satellites: |
0 |
0 bp |
0.00% |
||
|
Simple repeats: |
0 |
0 bp |
0.00% |
|
Simple repeats: |
0 |
0 bp |
0.00% |
||
|
Low complexity: |
0 |
0 bp |
0.00% |
|
Low complexity: |
0 |
0 bp |
0.00% |
||
D. Mito-genome Map
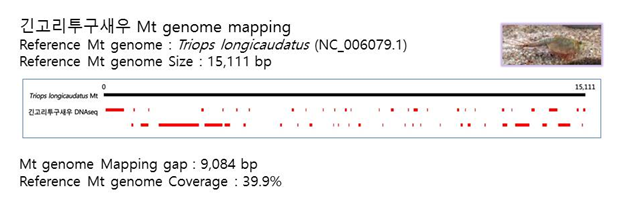
(4) Microsatellite 후보군
A. SSR statistics
|
Motif(-mer) |
2 |
3 |
4 |
5 |
6 |
7 |
8 |
9 |
10 |
Total |
|
긴꼬리투구새우 |
3,461 |
1,585 |
117 |
7 |
21 |
1 |
- |
- |
- |
5,192 |
B. 바코드서열
>긴꼬리투구새우(Triops_longicaudatus)
GAGCTGGGATAGTAGGAACAGCTCTGAGTCTTTTTATTCGAGCAGAAGTAGGCCAACCAA
GAAGACTGATCGGAGATGACCAAATTTACAACGTAATTGTAACAGCCCATGCATTTATTA
TAAGATTTTTTATGGTTATACCTATTTTAATTGGGGGTTTGGTAATTGGCTAGTACCATT
AATATTAGGGGCCCCTGACATGGCCTTCCCTCGTTTATATAATATAAGATTCTGATTATT
ACCTCCAGCCCTTACTTTATTATTATCAGGAGGTGCTGTAGAAGGAGGAGCTGGGACAGG
TTGAACAGTTTATCCTCCTCTATCAGGAGGAACTGCTCATGCGGGAACATTAGTAAACTG
AGGTATATTTTCATTACATTTAGCTGGGTTTCACCAATTTTAGGTGCTATTTATTTTATC
ACAGCAATCATTATTTACGAACCAGAGACATGTCTTTAGATCGAATCCCATTATTCGTTC
GAGCCGTAGGGATCACCACTTTACTTCTCGTCTCATTTCCTGTTCTAGCAGGGGGCAATC
ACAATATTATTAACGGACCGAAAGTTAAATACATCATTCTTCGACCCAGCTGGAGGAGGG
GATCCAATTCTTTATCAACATCTTTTCTGATTTTTTGG