쌍꼬리부전나비 (Spindasis takanonis)
|
|
▣ 국 명: 쌍꼬리부전나비 ▣ 학 명: Spindasis takanonis ▣ 지정번호: 멸종위기동물II급 ▣ 계 통: 곤충강-나비목-부전나비과 |
◎ 생김새 : 암수 모두 전체 몸길이는 28~40mm이고 날개 길이는 13~18mm 내외이다. 날개 앞면은 흑갈색이고 수컷의 경우 자줏빛이 도는 보라색의 광택을 보인다. 수컷은 암컷보다 약간 작고, 앞날개 바깥 측면의 곡면이 암컷보다 직선형으로 나타난다. 날개 앞면은 암갈색으로 수컷은 자줏빛 광택을 띠고 있다. 뒷날개의 후각은 주황색으로 검은 점이 속에 있고, 1쌍의 미상돌기를 갖는다. 날개를 폈을 때 뒷면은 담황색에 검정 줄무늬나 점무늬가 선명한 줄이 형성하고 있다.
◎ 서식지 : 개밋과의 한 종인 마쓰무라꼬리치레개미가 주고 살고 있는 벚나무나 소나무 등 고목, 그 일부분이 고사되어 이들이 정착한 곳에 주로 서식처를 만든다.
◎ 먹이 : 개망초 등의 꽃에서 꿀을 빨아먹으며, 애벌레 시절에는 숙주와의 먹이교환 행동을 통하거나, 나무속에 들어가 개미집을 찾아 개미 집안의 저장 먹이를 이용한다. 숙주 개미집 속에서 애벌레는 직접 먹이를 훔쳐 먹거나 구걸 행동 후 개미와 서로 입을 맞대고 구토 물질을 받아먹기도 한다.
◎ 번식 : 알 낳는 시기는 6월 중순부터 7월 초까지이며, 오후 5~6시 무렵에 알을 낳는다. 암컷은 숙주 개미인 마쓰무리밀드리개미의 집이 있는 소나무와 신갈나무?노간주나무나 바위틈에 알을 낳는데, 나무가 죽어 있거나 살아 있거나 상관하지 않는다. 한 번에 1~3개씩 같은 장소에 알을 낳기도 한다. 알을 낳은 지 3일 뒤 애벌레가 깨어나며 개미집과 떨어져 있는 경우 그곳을 지나는 숙주 일개미에 의해 옮겨지고 집 입구의 경우 애벌레가 직접 그 안으로 들어간다.
◎ 분포 : 한국?일본?중국 서부에 분포하며 한반도에서는 국지적 분포를 보인다.
1) 채집정보
- 채집허가: 2014.07.17 / 5마리
- 채 집 자: 빛고을생태연구소 정헌천
2) 채집현황 및 샘플처리
|
쌍꼬리부전나비 채집현황 |
샘플처리 |
|||
|
채집일 |
개체 수 |
DNAseq |
RNAseq |
표본 |
|
2014.07.30 |
5 EA |
3 EA |
2 EA |
- |
3) QC Data
|
샘플종 |
DNA QC |
RNA QC |
Result |
||
|
Concentration (ug/ul) |
Total Conc. (ug) |
Concentration (ug/ul) |
RIN (>7) |
||
|
쌍꼬리부전나비 |
64 |
12.8 |
306 |
8.9 |
pass |
4) 기존 NCBI Taxonomy 유전정보 현황
|
유전정보 종류 |
Spindasis 속 유전자원 |
Spindasis takanonis 유전자원 |
|
Nucleotide |
21 |
9 |
|
Protein |
41 |
33 |
|
Genome |
1 |
1 |
|
Gene |
13 |
13 |
5) Sequencing 분석 결과
- Illumina DNA Sequncing 결과
|
Input information |
|||||
|
|
|
|
Region1 |
Region2 |
Total |
|
raw data |
|||||
|
Number Of Reads |
|
|
134,720,924 |
134,720,924 |
269,441,848 |
|
Number Of Bases |
|
|
16,974,836,424 |
16,974,836,424 |
33,949,672,848 |
|
error correction |
|||||
|
Number Of Reads |
|
|
133,867,320 |
133,007,479 |
266,874,799 |
|
Number Of Bases |
|
|
16,681,597,553 |
16,491,539,010 |
33,173,136,563 |
|
pair reads |
|||||
|
Number Of Reads |
|
|
|
|
265,227,508 |
|
Number Of Bases |
|
|
|
|
32,979,377,831 |
|
single reads |
|||||
|
Number Of Reads |
|
|
|
|
1,647,291 |
|
Number Of Bases |
|
|
|
|
193,758,732 |
|
Assembly Results |
|||||
|
Scaffold Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Scaffolds |
285,846 |
285,846 |
177,218 |
132,257 |
81,403 |
|
Number Of Bases |
501,954,621 |
501,954,621 |
478,363,926 |
445,636,909 |
372,137,312 |
|
Avg. Scaffold Size |
1,756 |
1,756 |
2,699 |
3,369 |
4,571 |
|
N50 Scaffold Size |
3,820 |
3,820 |
4,033 |
4,347 |
5,115 |
|
N80 Scaffold Size |
1,600 |
1,600 |
1,853 |
2,213 |
3,058 |
|
N90 Scaffold Size |
911 |
911 |
1,211 |
1,607 |
2,514 |
|
largest Scaffold Size |
130,357 |
130,357 |
130,357 |
130,357 |
130,357 |
|
Contig Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Contigs |
8,871,600 |
959,759 |
317,010 |
131,954 |
32,715 |
|
Number Of Bases |
833,507,629 |
502,450,437 |
361,672,343 |
231,092,953 |
95,481,504 |
|
Avg. Contig Size |
93 |
523 |
1,140 |
1,751 |
2,918 |
|
N50 Contig Size |
318 |
907 |
1,270 |
1,776 |
2,830 |
|
N80 Contig Size |
38 |
367 |
753 |
1,246 |
2,262 |
|
N90 Contig Size |
33 |
204 |
621 |
1,115 |
2,123 |
|
Largest Contig Size |
45,422 |
45,422 |
45,422 |
45,422 |
45,422 |
- Illumina RNA Sequncing 결과
|
Input information |
|||||
|
|
|
|
Region1 |
Region2 |
Total |
|
raw data |
|||||
|
Number Of Reads |
|
|
124,656,396 |
124,656,396 |
249,312,792 |
|
Number Of Bases |
|
|
15,706,705,896 |
15,706,705,896 |
31,413,411,792 |
|
error correction |
|||||
|
Number Of Reads |
|
|
123,652,370 |
122,810,483 |
246,462,853 |
|
Number Of Bases |
|
|
15,376,471,577 |
15,259,523,350 |
30,635,994,927 |
|
pair reads |
|||||
|
Number Of Reads |
|
|
|
|
244,865,978 |
|
Number Of Bases |
|
|
|
|
30,453,770,932 |
|
single reads |
|||||
|
Number Of Reads |
|
|
|
|
1,596,875 |
|
Number Of Bases |
|
|
|
|
182,223,995 |
|
Assembly Results |
|||||
|
Scaffold Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Scaffolds |
94,820 |
94,820 |
23,344 |
10,607 |
3,737 |
|
Number Of Bases |
45,575,743 |
45,575,743 |
30,312,925 |
21,439,648 |
11,881,542 |
|
Avg. Scaffold Size |
480 |
480 |
1,298 |
2,021 |
3,179 |
|
N50 Scaffold Size |
903 |
903 |
1,588 |
2,179 |
3,205 |
|
N80 Scaffold Size |
288 |
288 |
808 |
1,373 |
2,384 |
|
N90 Scaffold Size |
176 |
176 |
637 |
1,169 |
2,182 |
|
largest Scaffold Size |
22,120 |
22,120 |
22,120 |
22,120 |
22,120 |
|
Contig Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Contigs |
1,533,314 |
137,430 |
22,873 |
7,765 |
1,501 |
|
Number Of Bases |
110,785,207 |
46,236,487 |
22,935,288 |
12,535,086 |
4,059,412 |
|
Avg. Contig Size |
72 |
336 |
1,002 |
1,614 |
2,704 |
|
N50 Contig Size |
63 |
493 |
1,078 |
1,613 |
2,607 |
|
N80 Contig Size |
38 |
193 |
671 |
1,197 |
2,195 |
|
N90 Contig Size |
32 |
139 |
580 |
1,092 |
2,091 |
|
Largest Contig Size |
21,743 |
21,743 |
21,743 |
21,743 |
21,743 |
(1) 고품질 genomic DNA 채취 및 표본확보
|
검수용 샘플 DNA 농도 및 총 양, 확증표본 현황표 |
|||||||
|
Sample명 |
농도(ng/ul) |
Volume(ul) |
DNA 총량(ug) |
O.D A260/280 |
확증표본 |
채집허가일 |
채집일 |
|
쌍고리부전나비 |
31 |
996 |
30 |
2.01 |
○ |
2014.7.17 |
2014.7.30 |
(2) 유전체 서열확보 : 현재 20종 확보
|
Sample |
구분 |
raw data size |
trim data size |
비고 |
|
쌍고리부전나비 |
DNA |
33,949,672,848 |
33,173,136,563 |
|
|
RNA |
31,413,411,792 |
30,635,994,927 |
|
(3) 기타분석
A. Genome Size estimation
|
샘 플 명 |
Kmer size |
Kmer total |
Pk depth |
Genome_size |
Coverage(X) |
|
쌍꼬리부전나비 |
23 |
21,016,464,144 |
29 |
724,705,660 |
37.1795 |
B. 반복서열
- 시퀀싱이 완료된 쌍꼬리부전나비 DNA 및 RNA의 contigs 중 1kb 이상의 contigs의 반복서열을 탐색(Left; Repeat masking results of DNA contigs, Right; Repeat masking results of RNA contigs).
|
sequences: 131,954 |
|
sequences: 7,765 |
||||||||
|
total length: 231,092,953 bp (231,092,953 bp excl N/X-runs) |
|
totallength:12,535,086bp (12,535,086 bp excl N/X-runs) |
||||||||
|
GC level: 33.94 % |
|
GC level: 39.24 % |
||||||||
|
bases masked: 47,786 bp ( 0.02 %) |
|
bases masked: 4,123 bp ( 0.03 %) |
||||||||
|
|
numberof elements |
length occupied |
percentage of sequence |
|
|
numberof elements |
length occupied |
percentage of sequence |
||
|
SINEs: |
68 |
4152 bp |
0.00% |
|
SINEs: |
68 |
4152 bp |
0.00% |
||
|
|
ALUs |
0 |
0 bp |
0.00% |
|
|
ALUs |
0 |
0 bp |
0.00% |
|
|
MIRs |
58 |
3641 bp |
0.00% |
|
|
MIRs |
58 |
3641 bp |
0.00% |
|
LINEs: |
105 |
6517 bp |
0.00% |
|
LINEs: |
105 |
6517 bp |
0.00% |
||
|
|
LINE1 |
16 |
839 bp |
0.00% |
|
|
LINE1 |
16 |
839 bp |
0.00% |
|
|
LINE2 |
15 |
830 bp |
0.00% |
|
|
LINE2 |
15 |
830 bp |
0.00% |
|
|
L3/CR1 |
50 |
3347 bp |
0.00% |
|
|
L3/CR1 |
50 |
3347 bp |
0.00% |
|
LTRelements: |
26 |
1667 bp |
0.00% |
|
LTRelements: |
26 |
1667 bp |
0.00% |
||
|
|
ERVL |
2 |
178 bp |
0.00% |
|
|
ERVL |
2 |
178 bp |
0.00% |
|
|
ERVL-MaLRs |
4 |
253 bp |
0.00% |
|
|
ERVL-MaLRs |
4 |
253 bp |
0.00% |
|
|
ERV_classI |
14 |
870 bp |
0.00% |
|
|
ERV_classI |
14 |
870 bp |
0.00% |
|
|
ERV_classII |
2 |
152 bp |
0.00% |
|
|
ERV_classII |
2 |
152 bp |
0.00% |
|
DNAelements: |
206 |
24819 bp |
0.01% |
|
DNAelements: |
206 |
24819 bp |
0.01% |
||
|
|
hAT-Charlie |
59 |
6672 bp |
0.00% |
|
|
hAT-Charlie |
59 |
6672 bp |
0.00% |
|
|
TcMar-Tigger |
24 |
2013 bp |
0.00% |
|
|
TcMar-Tigger |
24 |
2013 bp |
0.00% |
|
Unclassified: |
11 |
742 bp |
0.00% |
|
Unclassified: |
11 |
742 bp |
0.00% |
||
|
Total interspersed repeats: |
|
37897 bp |
0.02% |
|
Total interspersed repeats: |
|
37897 bp |
0.02% |
||
|
Small RNA: |
183 |
9959 bp |
0.00% |
|
Small RNA: |
183 |
9959 bp |
0.00% |
||
|
Satellites: |
1 |
101 bp |
0.00% |
|
Satellites: |
1 |
101 bp |
0.00% |
||
|
Simple repeats: |
0 |
0 bp |
0.00% |
|
Simple repeats: |
0 |
0 bp |
0.00% |
||
|
Low complexity: |
0 |
0 bp |
0.00% |
|
Low complexity: |
0 |
0 bp |
0.00% |
||
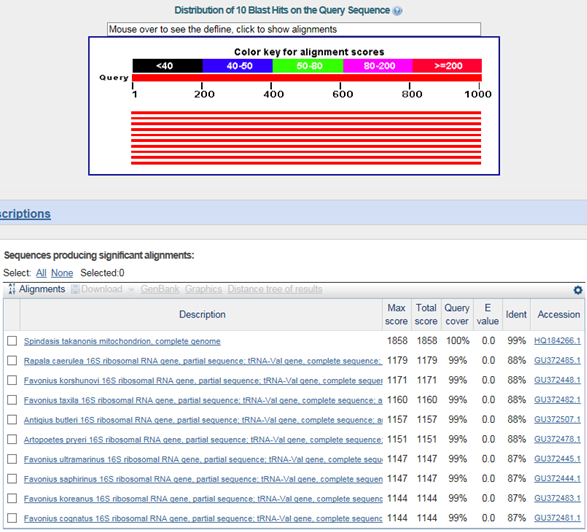
D. Mito-genome Map
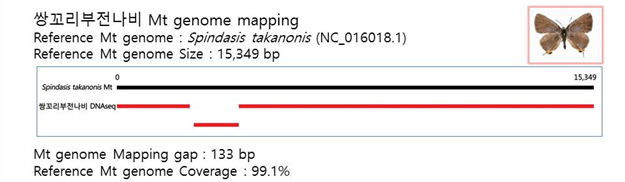
(4) Microsatellite 후보군
A. SSR statistics
|
Motif(-mer) |
2 |
3 |
4 |
5 |
6 |
7 |
8 |
9 |
10 |
Total |
|
쌍꼬리부전나비 |
17,853 |
3,747 |
1,131 |
157 |
30 |
1 |
- |
- |
- |
22,919 |
B. 바코드서열
>쌍꼬리부전나비(Spindasis takanonis)
AAAAAAAAGTTTATTTAATTTTTTTATCACCCCAATAAAATAATTAATAAATTTAAAATT
TAAAAATTTATATAAAATTTTAAATTTATTAATTATAAAACTCTATAGGGTCTTCTCGTC
TTTTAAATATATTTTAACTTTTTAATTAAAAAATTAAATTCTAAAATTAATAATGAGACA
GTATATATTTCATCAAATCTTTCATACAAGTCTTCAATTAAAAGACTAATGATTATGCTA
CCTTTGTACAGTCAAAATACTGCAGCCCTTTAATAATTATCAGTGGGCAGATTAGACTTT
AAATTATTATATCAAAAAGACATGTTTTTGTTAAACAGGTGAATATAAATTTTTGCCGAA
TTCCTTTTTATTATTTATTAAAAAATTAATAAATTTCTTTATTTTTATACTAATTTTATC
ATTATAACAATTACTTAAAAATTAAATAATATTTTTTTATAAAAAATTAAATTAAATTAA
ATTTTACTTATTTTAAAATATTTTATAATAAATTTTAATTTTTAAACAATTTAAATTTAA
AAAATTTTAAAATACTTATCTAATTTATTATAAAAATTTAATTTAAAGCTTATCCCTTAA
ATTATTAAATTTAAAAATATAAAAATTAAAAATATAATTTTTATAAATATATAAATTTAA
AATTAAATTTTTTTCTAAAAAAACTAGATAATTTTAAAAACGATTAACATTTCATTTCAA
ATTAATTATTTAAAATAATTATATCACATTAAATTTATTAATTAATTAACTCTTTTAAAT
TCGAGATTTTTTAAAAATAAAAATTTAATTAATAAACTCTGATACACAAGATACATTAAA
TTAAAACTACTTTTATAAAAATTTATTTTCACATATTTCAAATTTCTTTTACAATACTAA
TTCCCTATAAATATAATAATTTTTTCTATTAAAATAAAATAACCCCCATTATTTATTTTA
TATTAAAATTATTTTTATTATAAATTTATTTTTATTAATTTTTATTTTA