붉은발말똥게 (Sesarma intermedium)
|
|
▣ 국명: 붉은발말똥게 ▣ 학명: Sesarma intermedium ▣ 지정번호: 멸종위기야생동물 Ⅱ급 ▣ 계통: 십각 목, Sesarmidae 과 |
1) 채집정보
- 채집허가: 2014.07.10 / 5마리
- 채 집 자: 신라대학교 고현숙
2) 채집현황 및 샘플처리
|
붉은발말똥게 채집현황 |
샘플처리 |
|||
|
채집일 |
개체 수 |
DNAseq |
RNAseq |
표본 |
|
2014.09.29 |
5 EA |
3 EA |
2 EA |
- |
3) QC Data
|
샘플종 |
DNA QC |
RNA QC |
Result |
||
|
Concentration (ug/ul) |
Total Conc. (ug) |
Concentration (ug/ul) |
RIN (>7) |
||
|
붉은발말똥게 |
108.4 |
20 |
353 |
6.3 |
pass |
4) 기존 NCBI Taxonomy 유전정보 현황
|
유전정보 종류 |
Sesarma 속 유전자원 |
Sesarma intermedium 유전자원 |
|
Nucleotide |
716 |
- |
|
Protein |
161 |
- |
|
Genome |
1 |
- |
|
Gene |
37 |
- |
5) Sequencing 분석 결과
- Illumina DNA Sequncing 결과
|
Input information |
|||||
|
|
|
|
Region1 |
Region2 |
Total |
|
raw data |
|||||
|
Number Of Reads |
|
|
189,843,243 |
189,843,243 |
379,686,486 |
|
Number Of Bases |
|
|
23,920,248,618 |
23,920,248,618 |
47,840,497,236 |
|
error correction |
|||||
|
Number Of Reads |
|
|
168,664,439 |
165,363,821 |
334,028,260 |
|
Number Of Bases |
|
|
18,538,231,785 |
18,135,415,500 |
36,673,647,285 |
|
pair reads |
|||||
|
Number Of Reads |
|
|
|
|
308,096,906 |
|
Number Of Bases |
|
|
|
|
34,351,794,449 |
|
single reads |
|||||
|
Number Of Reads |
|
|
|
|
25,931,354 |
|
Number Of Bases |
|
|
|
|
2,321,852,836 |
|
Assembly Results |
|||||
|
Scaffold Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Scaffolds |
3,203,608 |
3,203,608 |
318,015 |
72,201 |
10,641 |
|
Number Of Bases |
843,259,159 |
843,259,159 |
276,146,740 |
110,831,892 |
29,880,453 |
|
Avg. Scaffold Size |
263 |
263 |
868 |
1,535 |
2,808 |
|
N50 Scaffold Size |
321 |
321 |
864 |
1,484 |
2,683 |
|
N80 Scaffold Size |
157 |
157 |
607 |
1,146 |
2,202 |
|
N90 Scaffold Size |
126 |
126 |
549 |
1,069 |
2,093 |
|
largest Scaffold Size |
259,778 |
259,778 |
259,778 |
259,778 |
259,778 |
|
Contig Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Contigs |
42,215,912 |
4,464,853 |
147,548 |
17,176 |
1,266 |
|
Number Of Bases |
3,021,343,702 |
892,697,805 |
107,928,053 |
23,583,775 |
3,519,700 |
|
Avg. Contig Size |
71 |
199 |
731 |
1,373 |
2,780 |
|
N50 Contig Size |
65 |
205 |
700 |
1,301 |
2,501 |
|
N80 Contig Size |
48 |
139 |
559 |
1,090 |
2,153 |
|
N90 Contig Size |
46 |
116 |
527 |
1,041 |
2,072 |
|
Largest Contig Size |
45,085 |
45,085 |
45,085 |
45,085 |
45,085 |
- Illumina RNA Sequncing 결과
|
Input information |
|||||
|
|
|
|
Region1 |
Region2 |
Total |
|
raw data |
|||||
|
Number Of Reads |
|
|
148,607,137 |
148,607,137 |
297,214,274 |
|
Number Of Bases |
|
|
18,724,499,262 |
18,724,499,262 |
37,448,998,524 |
|
error correction |
|||||
|
Number Of Reads |
|
|
144,869,389 |
143,377,719 |
289,738,778 |
|
Number Of Bases |
|
|
17,651,509,949 |
17,423,679,549 |
35,075,189,498 |
|
pair reads |
|||||
|
Number Of Reads |
|
|
|
|
284,329,580 |
|
Number Of Bases |
|
|
|
|
34,683,772,704 |
|
single reads |
|||||
|
Number Of Reads |
|
|
|
|
3,917,528 |
|
Number Of Bases |
|
|
|
|
391,416,794 |
|
Assembly Results |
|||||
|
Scaffold Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Scaffolds |
429,442 |
429,442 |
37,082 |
11,095 |
2,764 |
|
Number Of Bases |
108,908,082 |
108,908,082 |
37,496,303 |
19,934,251 |
8,708,719 |
|
Avg. Scaffold Size |
253 |
253 |
1,011 |
1,796 |
3,150 |
|
N50 Scaffold Size |
302 |
302 |
1,060 |
1,815 |
3,094 |
|
N80 Scaffold Size |
145 |
145 |
648 |
1,232 |
2,333 |
|
N90 Scaffold Size |
119 |
119 |
566 |
1,106 |
2,157 |
|
largest Scaffold Size |
15,520 |
15,520 |
15,520 |
15,520 |
15,520 |
|
Contig Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Contigs |
6,071,178 |
559,812 |
25,602 |
6,322 |
1,109 |
|
Number Of Bases |
449,202,856 |
113,481,169 |
22,996,315 |
10,066,265 |
3,115,851 |
|
Avg. Contig Size |
73 |
202 |
898 |
1,592 |
2,809 |
|
N50 Contig Size |
74 |
202 |
904 |
1,570 |
2,710 |
|
N80 Contig Size |
49 |
130 |
612 |
1,172 |
2,207 |
|
N90 Contig Size |
46 |
113 |
551 |
1,078 |
2,086 |
|
Largest Contig Size |
11,458 |
11,458 |
11,458 |
11,458 |
11,458 |
(1) 고품질 genomic DNA 채취 및 표본확보
|
검수용 샘플 DNA 농도 및 총 양, 확증표본 현황표 |
|||||||
|
Sample명 |
농도(ng/ul) |
Volume(ul) |
DNA 총량(ug) |
O.D A260/280 |
확증표본 |
채집허가일 |
채집일 |
|
붉은발말똥게 |
211 |
996 |
210 |
2.16 |
X |
2014.7.10 |
2014.9.29 |
(2) 유전체 서열확보 : 현재 20종 확보
|
Sample |
구분 |
raw data size |
trim data size |
비고 |
|
붉은발말똥게 |
DNA |
47,840,497,236 |
36,673,647,285 |
|
|
RNA |
37,448,998,524 |
35,075,189,498 |
|
(3) 기타분석
A. Genome Size estimation
|
샘 플 명 |
Kmer size |
Kmer total |
Pk depth |
Genome_size |
Coverage(X) |
|
붉은발말똥게 |
23 |
29,615,545,908 |
202 |
146,611,613 |
258.974 |
B. 반복서열
- 시퀀싱이 완료된 붉은발말똥게 DNA 및 RNA의 contigs 중 1kb 이상의 contigs의 반복서열을 탐색(Left; Repeat masking results of DNA contigs, Right; Repeat masking results of RNA contigs).
|
sequences: 17,176 |
|
sequences: 6,322 |
||||||||
|
total length: 23,583,775 bp (23,583,775 bp excl N/X-runs) |
|
total length: 10,066,265 bp (10,066,265 bp excl N/X-runs) |
||||||||
|
GC level: 39.60% |
|
GC level: 48.04% |
||||||||
|
bases masked: 87,632 bp (0.37%) |
|
bases masked: 15,587 bp (0.15%) |
||||||||
|
|
number of elements |
length occupied |
percentage of sequence |
|
|
number of elements |
length occupied |
percentage of sequence |
||
|
SINEs: |
5 |
302 bp |
0.00% |
|
SINEs: |
1 |
47 bp |
0.00% |
||
|
|
ALUs |
0 |
0 bp |
0.00% |
|
|
ALUs |
0 |
0 bp |
0.00% |
|
|
MIRs |
5 |
302 bp |
0.00% |
|
|
MIRs |
1 |
47 bp |
0.00% |
|
LINEs: |
218 |
38,074 bp |
0.16% |
|
LINEs: |
34 |
4,266 bp |
0.04% |
||
|
|
LINE1 |
6 |
272 bp |
0.00% |
|
|
LINE1 |
3 |
130 bp |
0.00% |
|
|
LINE2 |
5 |
363 bp |
0.00% |
|
|
LINE2 |
7 |
425 bp |
0.00% |
|
|
L3/CR1 |
186 |
31,756 bp |
0.13% |
|
|
L3/CR1 |
18 |
2,425 bp |
0.02% |
|
LTR elements: |
13 |
1,559 bp |
0.01% |
|
LTR elements: |
8 |
1,758 bp |
0.02% |
||
|
|
ERVL |
2 |
147 bp |
0.00% |
|
|
ERVL |
0 |
0 bp |
0.00% |
|
|
ERVL-MaLRs |
1 |
67 bp |
0.00% |
|
|
ERVL-MaLRs |
0 |
0 bp |
0.00% |
|
|
ERV_classI |
7 |
565 bp |
0.00% |
|
|
ERV_classI |
4 |
226 bp |
0.00% |
|
|
ERV_classII |
0 |
0 bp |
0.00% |
|
|
ERV_classII |
0 |
0 bp |
0.00% |
|
DNA elements: |
257 |
41,922 bp |
0.18% |
|
DNA elements: |
46 |
7,968 bp |
0.08% |
||
|
|
hAT-Charlie |
89 |
19,292 bp |
0.08% |
|
|
hAT-Charlie |
10 |
2,972 bp |
0.03% |
|
|
TcMar-Tigger |
101 |
12,506 bp |
0.05% |
|
|
TcMar-Tigger |
22 |
3,080 bp |
0.03% |
|
Unclassified: |
0 |
0 bp |
0.00% |
|
Unclassified: |
0 |
0 bp |
0.00% |
||
|
Total interspersed repeats: |
|
81,857 bp |
0.35% |
|
Total interspersed repeats: |
|
14,039 bp |
0.14% |
||
|
Small RNA: |
109 |
5,775 bp |
0.02% |
|
Small RNA: |
18 |
1,548 bp |
0.02% |
||
|
Satellites: |
0 |
0 bp |
0.00% |
|
Satellites: |
0 |
0 bp |
0.00% |
||
|
Simple repeats: |
0 |
0 bp |
0.00% |
|
Simple repeats: |
0 |
0 bp |
0.00% |
||
|
Low complexity: |
0 |
0 bp |
0.00% |
|
Low complexity: |
0 |
0 bp |
0.00% |
||
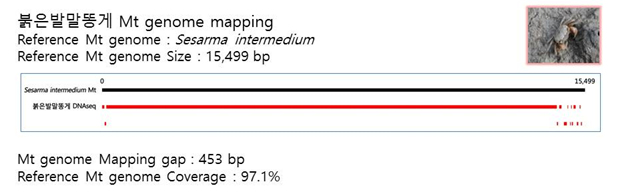
D. Mito-genome Map

(4) Microsatellite 후보군
A. SSR statistics
|
Motif(-mer) |
2 |
3 |
4 |
5 |
6 |
7 |
8 |
9 |
10 |
Total |
|
붉은발말똥개 |
3,778 |
1,699 |
330 |
34 |
- |
1 |
- |
- |
- |
5,842 |
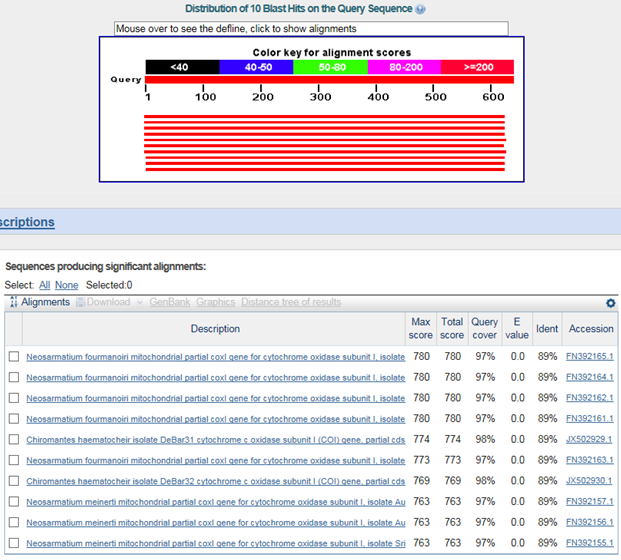
B. 바코드서열
>붉은발말똥게(Sesarma intermedium)
CCTGCAGGATTAAAGAAGGATGTATTTAAATTACGATCAATTAAAAGTATGGAAATAGGA
CCAGCTAAAACTGGAAGAGATAACAGTAATCAAATAGCATAAATAAATACTGATCATACA
AATAAAGGTATTTGATGTATAGTTATACCATAAGATCGTATTTTAATTACTGTTGTTATA
AAATTTATTGCTCCTCAAATAGATGAAACACCAGCTAGATGTCAAGAAAAAAATCCTAAA
TCTACAGAGGCCCCTGCATGAGCAATAGCAGCCGCTAAAGGAGGATACACAGTTCATCCT
GTTCCAACACCTCTTTCAACTATGCTTCTTGTTAATAATAATGATAAAGAAGGAGGTAGC
AATCAAAATCTTATATTATTTATACGTGGAAATGCCATATCTGGAGCTCCTAGTATTAAG
GGAACTAATCAATTGCCAAACCCTCCAATTATAATAGGAATAACTATAAAAAAAAATTAT
TACAAATGCATGGGCTGTTACAATTACATTATAAATTTGATCATTACCAATTAATCTTCC
TGGTTCATTTAATTCTGCTCGAATAATTAATCTTAATGATATACCTACTATTCCTGCTCA
AGCCAAAAAAACAAATATAAAGTGCCAATATCTTT
Neosarmatium fourmanoiri : family 까지 같음
chiromantes haematocheir : family 까지 같음