АХСІПмСйДоЦиРЬ (Satsuma myomphala)
|
|
ЂУ БЙ Иэ: АХСІПмСйДоЦиРЬ ЂУ Ча Иэ: Satsuma myomphala ЂУ СіСЄЙјШЃ: АќТћСО ЂУ Аш Хы: КЙСЗА-СјРЏЦѓИё-ПмСйДоЦиРЬАњ |
Ён Л§БшЛѕ : ЦаАЂЧќХТДТ ГЊХОРЬ ГЗРК ПјУпЧќРИЗЮ ГЊУўРК 6.5УўРЬДй. ЦаАЂРЧ АЂЧЧДТ ШВАЅЛіРЬАэ УМУўРЬ ХЉАэ ЕеБлДй. УМУў СпОгПЁМ НУРлЧи ТїУМСѕРЧ КРЧе РЇЗЮ РЬОюСіДТ РћАЅЛі ЛіДыАЁ РжДй КРЧеРК ОшАэ АЁДТ МКРхИЦРЬ КёНКЕыШї РжРИИч СІАјРК СМАэ УрМј БйЙцПЁМ ДнШљДй. АЂБИДТ ЙнПјЧќРИЗЮ ГаАэ УрМј КЮРЇАЁ СЅЧєСјДй. МКУМРЧ ХЉБтДТ ВЎЕЅБт ГєРЬ 27mm, СіИЇ 37mmСЄЕЕРЬДй.
Ён МНФСі : ЙйДйАЁ РЮСЂЧб НРЧЯАэ ПьАХСј ГДыИВ НЃМгПЁ МНФЧбДй.
Ён ПЌБИРЧ ЧЪПфМК : АГУМБКРЧ ХЉБтАЁ ИХПь РлАэ, УтЧіЙќРЇЕЕ ИХПь ЧљМвЧЯДй. ДыЧќСОРИЗЮ УЕРћПЁ РЧЧи ЦїНФДчЧЯАХГЊ ПьБтПЁ НЃРЛ ЙўОюГ АГУМРЧ АцПь ОаЛч ЖЧДТ ГВШЙРЧ ПьЗСАЁ ГєРИИч МНФУГРЧ АГЙп Йз СњРЧ ЧЯЖєРЬ АЁРх ХЋ РЇЧљ ПфРЮРЬДй.
Ён КаЦї : ЧбБЙ, РЯКЛПЁ КаЦїЧЯАэ РжРИИч ЧбБЙРЧ АцПь АцЛѓГВЕЕ АХСІНУ АЅАљИЎРЧ СМРК ЙќРЇ ГЛПЁМИИ ЙпАпЕШДй.
1) ӪѧѪКИ
- УЄС§ЧуАЁ: (АќТћСО)
- УЄ С§ Рк: АПјДыЧаБГ РЬСиЛѓ
2) УЄС§ЧіШВ Йз ЛљЧУУГИЎ
|
АХСІПмСйДоЦиРЬ УЄС§ЧіШВ |
ЛљЧУУГИЎ |
|||
|
УЄС§РЯ |
АГУМ Мі |
DNAseq |
RNAseq |
ЧЅКЛ |
|
2014.08.18 |
1 EA |
1 EA (Йп) |
1 EA (ГЛРх) |
- |
3) QC Data
|
ЛљЧУСО |
DNA QC |
RNA QC |
Result |
||
|
Concentration (ug/ul) |
Total Conc. (ug) |
Concentration (ug/ul) |
RIN (>7) |
||
|
АХСІПмСйДоЦиРЬ |
109 |
11.9 |
275 |
5.1 |
pass |
4) БтСИ NCBI Taxonomy РЏРќСЄКИ ЧіШВ
|
РЏРќСЄКИ СОЗљ |
Satsuma Мг РЏРќРкПј |
Satsuma myomphala РЏРќРкПј |
|
Nucleotide |
959 |
6 |
|
Protein |
522 |
2 |
5) Sequencing КаМЎ АсАњ
- Illumina DNA Sequncing АсАњ
|
Input information |
|||||
|
ЁЁ |
ЁЁ |
ЁЁ |
Region1 |
Region2 |
Total |
|
raw data |
|||||
|
Number Of Reads |
|
|
474,497,656 |
474,497,656 |
948,995,312 |
|
Number Of Bases |
|
|
59,786,704,656 |
59,786,704,656 |
119,573,409,312 |
|
error correction |
|||||
|
Number Of Reads |
|
|
472,040,161 |
463,548,129 |
935,588,290 |
|
Number Of Bases |
|
|
58,296,217,558 |
56,830,286,536 |
115,126,504,094 |
|
pair reads |
|||||
|
Number Of Reads |
|
|
|
|
922,815,600 |
|
Number Of Bases |
|
|
|
|
113,602,679,970 |
|
single reads |
|||||
|
Number Of Reads |
|
|
|
|
12,772,690 |
|
Number Of Bases |
|
|
|
|
1,523,824,124 |
|
Assembly Results |
|||||
|
Scaffold Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Scaffolds |
3,960,072 |
3,960,072 |
1,092,228 |
708,600 |
358,871 |
|
Number Of Bases |
2,628,937,244 |
2,628,937,244 |
2,138,912,269 |
1,863,107,832 |
1,363,667,228 |
|
Avg. Scaffold Size |
663 |
663 |
1,958 |
2,629 |
3,799 |
|
N50 Scaffold Size |
2,110 |
2,110 |
2,716 |
3,110 |
3,984 |
|
N80 Scaffold Size |
563 |
563 |
1,287 |
1,727 |
2,659 |
|
N90 Scaffold Size |
183 |
183 |
885 |
1,353 |
2,314 |
|
largest Scaffold Size |
35,616 |
35,616 |
35,616 |
35,616 |
35,616 |
|
Contig Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Contigs |
80,307,751 |
10,625,614 |
1,255,442 |
344,403 |
44,039 |
|
Number Of Bases |
6,740,211,639 |
2,891,139,666 |
1,137,687,242 |
511,274,653 |
114,388,576 |
|
Avg. Contig Size |
83 |
272 |
906 |
1,484 |
2,597 |
|
N50 Contig Size |
86 |
356 |
932 |
1,450 |
2,502 |
|
N80 Contig Size |
48 |
154 |
637 |
1,143 |
2,156 |
|
N90 Contig Size |
46 |
123 |
564 |
1,067 |
2,073 |
|
Largest Contig Size |
14,649 |
14,649 |
14,649 |
14,649 |
14,649 |
- Illumina RNA Sequncing АсАњ
|
Input information |
|||||
|
ЁЁ |
ЁЁ |
ЁЁ |
Region1 |
Region2 |
Total |
|
raw data |
|||||
|
Number Of Reads |
|
|
131,496,702 |
130,723,975 |
262,220,677 |
|
Number Of Bases |
|
|
16,388,085,286 |
16,308,351,332 |
32,696,436,618 |
|
error correction |
|||||
|
Number Of Reads |
|
|
129,324,855 |
128,180,861 |
257,505,716 |
|
Number Of Bases |
|
|
15,671,819,041 |
15,486,758,524 |
31,158,577,565 |
|
pair reads |
|||||
|
Number Of Reads |
|
|
|
|
260,815,568 |
|
Number Of Bases |
|
|
|
|
32,538,377,371 |
|
single reads |
|||||
|
Number Of Reads |
|
|
|
|
1,405,109 |
|
Number Of Bases |
|
|
|
|
158,059,247 |
|
Assembly Results |
|||||
|
Scaffold Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Scaffolds |
97,460 |
97,460 |
17,194 |
5,194 |
710 |
|
Number Of Bases |
32,605,181 |
32,605,181 |
16,073,800 |
7,810,353 |
1,805,352 |
|
Avg. Scaffold Size |
334 |
334 |
934 |
1,503 |
2,542 |
|
N50 Scaffold Size |
488 |
488 |
978 |
1,479 |
2,464 |
|
N80 Scaffold Size |
194 |
194 |
650 |
1,162 |
2,158 |
|
N90 Scaffold Size |
143 |
143 |
570 |
1,076 |
2,076 |
|
largest Scaffold Size |
9,386 |
9,386 |
9,386 |
9,386 |
9,386 |
|
Contig Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Contigs |
2,321,474 |
131,541 |
12,426 |
2,988 |
330 |
|
Number Of Bases |
128,115,613 |
32,754,540 |
10,714,009 |
4,317,505 |
813,111 |
|
Avg. Contig Size |
55 |
249 |
862 |
1,444 |
2,463 |
|
N50 Contig Size |
59 |
302 |
874 |
1,416 |
2,384 |
|
N80 Contig Size |
34 |
147 |
614 |
1,139 |
2,122 |
|
N90 Contig Size |
32 |
118 |
552 |
1,065 |
2,056 |
|
Largest Contig Size |
7,847 |
7,847 |
7,847 |
7,847 |
7,847 |
(1) АэЧАСњ genomic DNA УЄУы Йз ЧЅКЛШЎКИ
|
АЫМіПы ЛљЧУ DNA ГѓЕЕ Йз Уб Оч, ШЎСѕЧЅКЛ ЧіШВЧЅ |
|||||||
|
ЁЁSampleИэ |
ГѓЕЕ(ng/ul) |
Volume(ul) |
DNA УбЗЎ(ug) |
O.D A260/280 |
ШЎСѕЧЅКЛ |
УЄС§ЧуАЁРЯ |
УЄС§РЯ |
|
АХСІПмСйДоЦиРЬ |
499 |
996 |
497 |
1.76 |
X |
АэРЏСО |
2014.8.18 |
(2) РЏРќУМ МПШЎКИ : ЧіРч 20СО ШЎКИ
|
Sample |
БИКа |
raw data size |
trim data size |
КёАэ |
|
АХСІПмСйДоЦиРЬ |
DNA |
119,573,409,312 |
115,126,504,094 |
ЁЁ |
|
RNA |
33,320,993,328 |
32,696,436,618 |
ЁЁ |
(3) БтХИКаМЎ
A. Genome Size estimation
|
Лљ ЧУ Иэ |
Kmer size |
Kmer total |
Pk depth |
Genome_size |
Coverage(X) |
|
АХСІПмСйДоЦиРЬ |
23 |
74,021,634,336 |
12 |
6,168,469,528 |
15.3846 |
B. ЙнКЙМП
- НУФіНЬРЬ ПЯЗсЕШ АХСІПмСйДоЦиРЬ DNA Йз RNAРЧ contigs Сп 1kb РЬЛѓРЧ contigsРЧ ЙнКЙМПРЛ ХНЛі(Left; Repeat masking results of DNA contigs, Right; Repeat masking results of RNA contigs).
|
sequences: 344,403 |
|
sequences: 2,988 |
||||||||
|
total length: 511,274,653 bp (511,274,653 bp excl N/X-runs) |
|
total length: 4,317,505 bp (4,317,505 bp excl N/X-runs) |
||||||||
|
GC level: 39.67% |
|
GC level: 44.87% |
||||||||
|
bases masked: 16,345,250 bp (3.20%) |
|
bases masked: 1,671 bp (0.04%) |
||||||||
|
|
number of elements |
length occupied |
percentage of sequence |
|
|
number of elements |
length occupied |
percentage of sequence |
||
|
SINEs: |
75 |
4,989 bp |
0.00% |
|
SINEs: |
1 |
56 bp |
0.00% |
||
|
|
ALUs |
0 |
0 bp |
0.00% |
|
|
ALUs |
0 |
0 bp |
0.00% |
|
|
MIRs |
30 |
1640 bp |
0.00% |
|
|
MIRs |
0 |
0 bp |
0.00% |
|
LINEs: |
1,210 |
237,274 bp |
0.05% |
|
LINEs: |
11 |
955 bp |
0.02% |
||
|
|
LINE1 |
123 |
8,454 bp |
0.00% |
|
|
LINE1 |
2 |
145 bp |
0.00% |
|
|
LINE2 |
72 |
4,308 bp |
0.00% |
|
|
LINE2 |
1 |
64 bp |
0.00% |
|
|
L3/CR1 |
173 |
10,456 bp |
0.00% |
|
|
L3/CR1 |
3 |
160 bp |
0.00% |
|
LTR elements: |
743 |
130,316 bp |
0.03% |
|
LTR elements: |
2 |
110 bp |
0.00% |
||
|
|
ERVL |
14 |
889 bp |
0.00% |
|
|
ERVL |
0 |
0 bp |
0.00% |
|
|
ERVL-MaLRs |
9 |
564 bp |
0.00% |
|
|
ERVL-MaLRs |
0 |
0 bp |
0.00% |
|
|
ERV_classI |
497 |
92,952 bp |
0.02% |
|
|
ERV_classI |
1 |
62 bp |
0.00% |
|
|
ERV_classII |
7 |
344 bp |
0.00% |
|
|
ERV_classII |
0 |
0 bp |
0.00% |
|
DNA elements: |
3,628 |
420,405 bp |
0.08% |
|
DNA elements: |
5 |
405 bp |
0.01% |
||
|
|
hAT-Charlie |
1,319 |
175,367 bp |
0.03% |
|
|
hAT-Charlie |
2 |
176 bp |
0.00% |
|
|
TcMar-Tigger |
716 |
83,254 bp |
0.02% |
|
|
TcMar-Tigger |
0 |
0 bp |
0.00% |
|
Unclassified: |
16 |
1061 bp |
0.00% |
|
Unclassified: |
0 |
0 bp |
0.00% |
||
|
Total interspersed repeats: |
|
794,045 bp |
0.16% |
|
Total interspersed repeats: |
|
1,526 bp |
0.04% |
||
|
Small RNA: |
1429 |
166,350 bp |
0.03% |
|
Small RNA: |
1 |
145 bp |
0.00% |
||
|
Satellites: |
20 |
3,138 bp |
0.00% |
|
Satellites: |
0 |
0 bp |
0.00% |
||
|
Simple repeats: |
196,453 |
13,240,212 bp |
2.59% |
|
Simple repeats: |
0 |
0 bp |
0.00% |
||
|
Low complexity: |
21,124 |
2,145,969 bp |
0.42% |
|
Low complexity: |
0 |
0 bp |
0.00% |
||
D. Mito-genome Map
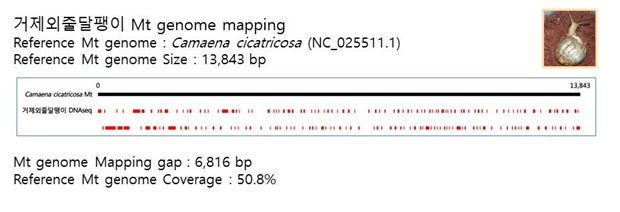
(4) Microsatellite ШФКИБК
A. SSR statistics
|
Motif(-mer) |
2 |
3 |
4 |
5 |
6 |
7 |
8 |
9 |
10 |
Total |
|
АХСІПмСйДоЦиРЬ |
119,557 |
52,648 |
15,821 |
1,121 |
91 |
38 |
32 |
16 |
6 |
189,330 |
B. ЙйФкЕхМП
>АХСІПмСйДоЦиРЬ(Satsuma myomphala)
ACTGTTAACAATGTAAAAATATAGATAATTAGGCTTATCTGAACTCGGCAAATTTTTCTT
CGACTGTTTATCAAAAACATAACCATTTGGTAGTATAAGTGGCTCTTTCTGCTCAATGAG
TAATTAAATAGCCGCAGTACCCTGACTGTGCAAAGGTAGCGTAATCATTTGGCTTTTAAT
TGGGGTCTAGTATGAACGAAAAATTGGGGAATAGCTTTCTCAGCTGAATAATTTGAATTT
TCTTAGTAGATGAAAATATCTACTCTTAATTAATAGACGAGAAGACCCTAGAAATTTTTA
TACAAGTTATTAGTATTTTTGTTGGGGCGACAGAGTAGCAACGAACAAACCTACTTGTTA
TTATTTTTCACCTATTTGGTATACAATTAAATTACTCTAGGGATAACAGCATAATGAAAT
TCGTTTGTGACCTCGATGTTGGACTAGGAAAACTTGGACCCAGAAGGTACATGTTAATGC
TCTGTTCGAGCAGATTCTCCTACATGATC