한국산개구리 (Rana coreana)
|
|
▣ 국 명: 한국산개구리 ▣ 영 명: Korean Brown Frog ▣ 학 명: Rana coreana ▣ 지정번호: 고유종 ▣ 계 통: 양서강-무미목-개구리과 |
1) 채집정보
- 채집허가: (고유종)
- 채 집 자: 서울대학교 민미숙
2) 채집현황 및 샘플처리
|
한국산개구리 채집현황 |
샘플처리 |
|||
|
채집일 |
개체 수 |
DNAseq |
RNAseq |
표본 |
|
2014.06.13 |
4 EA |
3 EA (몸통) |
3 EA (내장) |
1 EA, 70% EtOH |
3) QC Data
|
샘플종 |
DNA QC |
RNA QC |
Result |
||
|
Concentration (ug/ul) |
Total Conc. (ug) |
Concentration (ug/ul) |
RIN (>7) |
||
|
한국산개구리 |
204.5 |
36.8 |
941 |
7.9 |
pass |
4) 기존 NCBI Taxonomy 유전정보 현황
|
유전정보 종류 |
Rana 속 유전자원 |
Rana coreana 유전자원 |
|
Nucleotide |
623,872 |
46 |
|
Protein |
42,616 |
46 |
|
Genome |
13 |
- |
|
Gene |
682 |
- |
5) Sequencing 분석 결과
- Illumina DNA Sequncing 결과
|
Input information |
||||||||||
|
|
Region1 |
Region2 |
Region3 |
Region4 |
Region5 |
Region6 |
Total |
|||
|
raw data |
||||||||||
|
Number Of Reads |
50,575,870 |
50,575,870 |
51,296,117 |
51,296,117 |
24,836,362 |
24,836,362 |
253,416,698 |
|||
|
Number Of Bases |
6,372,559,620 |
6,372,559,620 |
6,463,310,742 |
6,463,310,742 |
3,129,381,612 |
3,129,381,612 |
31,930,503,948 |
|||
|
error correction |
||||||||||
|
Number Of Reads |
32,542,947 |
31,882,394 |
34,639,181 |
34,261,207 |
12,606,447 |
12,406,604 |
158,338,780 |
|||
|
Number Of Bases |
3,201,574,831 |
3,115,010,764 |
3,456,089,885 |
3,403,093,808 |
1,229,834,433 |
1,202,324,344 |
15,607,928,065 |
|||
|
pair reads |
||||||||||
|
Number Of Reads |
|
50,694,724 |
|
55,058,516 |
|
18,860,818 |
124,614,058 |
|||
|
Number Of Bases |
|
5,191,957,204 |
|
5,701,574,390 |
|
1,935,330,406 |
12,828,862,000 |
|||
|
single reads |
||||||||||
|
Number Of Reads |
|
13,730,617 |
|
13,841,872 |
|
6,152,233 |
33,724,722 |
|||
|
Number Of Bases |
|
1,124,628,391 |
|
1,157,609,303 |
|
496,828,371 |
2,779,066,065 |
|||
|
Assembly Results |
||||||||||
|
Scaffold Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|||||
|
Number Of Scaffolds |
762,316 |
762,316 |
7,716 |
556 |
39 |
|||||
|
Number Of Bases |
127,005,308 |
127,005,308 |
5,238,137 |
753,158 |
104,060 |
|||||
|
Avg. Scaffold Size |
166 |
166 |
678 |
1,354 |
2,668 |
|||||
|
N50 Scaffold Size |
162 |
162 |
641 |
1,273 |
2,501 |
|||||
|
N80 Scaffold Size |
122 |
122 |
541 |
1,085 |
2,166 |
|||||
|
N90 Scaffold Size |
108 |
108 |
519 |
1,038 |
2,073 |
|||||
|
largest Scaffold Size |
8,766 |
8,766 |
8,766 |
8,766 |
8,766 |
|||||
|
Contig Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|||||
|
Number Of Contigs |
22,666,825 |
844,997 |
2,566 |
223 |
23 |
|||||
|
Number Of Bases |
1,319,872,730 |
130,179,797 |
1,785,966 |
319,091 |
66,044 |
|||||
|
Avg. Contig Size |
58 |
154 |
696 |
1,430 |
2,871 |
|||||
|
N50 Contig Size |
53 |
153 |
650 |
1,320 |
2,536 |
|||||
|
N80 Contig Size |
47 |
119 |
542 |
1,109 |
2,238 |
|||||
|
N90 Contig Size |
46 |
108 |
519 |
1,057 |
2,157 |
|||||
|
Largest Contig Size |
8,766 |
8,766 |
8,766 |
8,766 |
8,766 |
|||||
- Illumina RNA Sequncing 결과
|
Input information |
|||||
|
|
|
|
Region1 |
Region2 |
Total |
|
raw data |
|||||
|
Number Of Reads |
|
|
119,908,635 |
119,908,635 |
239,817,270 |
|
Number Of Bases |
|
|
15,108,488,010 |
15,108,488,010 |
30,216,976,020 |
|
error correction |
|||||
|
Number Of Reads |
|
|
117,986,311 |
116,790,006 |
234,776,317 |
|
Number Of Bases |
|
|
14,611,372,750 |
14,434,014,702 |
29,045,387,452 |
|
pair reads |
|||||
|
Number Of Reads |
|
|
|
|
232,474,692 |
|
Number Of Bases |
|
|
|
|
28,795,593,860 |
|
single reads |
|||||
|
Number Of Reads |
|
|
|
|
2,301,625 |
|
Number Of Bases |
|
|
|
|
249,793,592 |
|
Assembly Results |
|||||
|
Scaffold Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Scaffolds |
168,341 |
168,341 |
32,466 |
13,150 |
4,024 |
|
Number Of Bases |
67,016,492 |
67,016,492 |
39,022,811 |
25,627,672 |
13,056,363 |
|
Avg. Scaffold Size |
398 |
398 |
1,201 |
1,948 |
3,244 |
|
N50 Scaffold Size |
666 |
666 |
1,390 |
2,028 |
3,210 |
|
N80 Scaffold Size |
226 |
226 |
749 |
1,317 |
2,376 |
|
N90 Scaffold Size |
152 |
152 |
613 |
1,149 |
2,163 |
|
largest Scaffold Size |
20,921 |
20,921 |
20,921 |
20,921 |
20,921 |
|
Contig Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Contigs |
2,832,511 |
237,434 |
27,131 |
8,481 |
1,677 |
|
Number Of Bases |
181,289,975 |
67,243,738 |
26,695,336 |
13,920,830 |
4,761,863 |
|
Avg. Contig Size |
64 |
283 |
983 |
1,641 |
2,839 |
|
N50 Contig Size |
63 |
364 |
1,037 |
1,626 |
2,737 |
|
N80 Contig Size |
35 |
162 |
656 |
1,197 |
2,234 |
|
N90 Contig Size |
32 |
126 |
572 |
1,089 |
2,114 |
|
Largest Contig Size |
13,557 |
13,557 |
13,557 |
13,557 |
13,557 |
|
Sample |
구분 |
raw data size |
trim data size |
비고 |
|
한국산개구리 |
DNA |
31,930,503,948 |
15,607,928,065 |
|
|
RNA |
30,216,976,020 |
29,045,387,452 |
|
(3) 기타분석
A. Genome Size estimation
|
샘 플 명 |
Kmer size |
Kmer total |
Pk depth |
Genome_size |
Coverage(X) |
|
한국산개구리 |
23 |
7,889,835,720 |
126 |
62,617,743 |
161.538 |
B. 반복서열
- 시퀀싱이 완료된 한국산개구리 DNA 및 RNA의 contigs 중 1kb 이상의 contigs의 반복서열을 탐색(Left; Repeat masking results of DNA contigs, Right; Repeat masking results of RNA contigs).
|
sequences: 132 |
|
sequences: 8481 |
||||||||
|
total length: 182,752 bp (182,752 bp excl N/X-runs) |
|
total length: 13,920,830 bp (13,920,830 bp excl N/X-runs) |
||||||||
|
GClevel: 43.18% |
|
GC level: 45.50% |
||||||||
|
bases masked: 5,851bp ( 3.20%) |
|
bases masked: 66,786 bp ( 0.48%) |
||||||||
|
|
numberof elements |
length occupied |
percentage of sequence |
|
|
numberof elements |
length occupied |
percentage of sequence |
||
|
SINEs: |
0 |
0 bp |
0.00% |
|
SINEs: |
0 |
0 bp |
0.00% |
||
|
|
ALUs |
0 |
0 bp |
0.00% |
|
|
ALUs |
0 |
0 bp |
0.00% |
|
|
MIRs |
0 |
0 bp |
0.00% |
|
|
MIRs |
0 |
0 bp |
0.00% |
|
LINEs: |
2 |
145 bp |
0.08% |
|
LINEs: |
17 |
1,906 bp |
0.01% |
||
|
|
LINE1 |
2 |
145 bp |
0.08% |
|
|
LINE1 |
7 |
1,195 bp |
0.01% |
|
|
LINE2 |
0 |
0 bp |
0.00% |
|
|
LINE2 |
2 |
132 bp |
0.00% |
|
|
L3/CR1 |
0 |
0 bp |
0.00% |
|
|
L3/CR1 |
6 |
488 bp |
0.00% |
|
LTRelements: |
1 |
714 bp |
0.39% |
|
LTRelements: |
5 |
563 bp |
0.00% |
||
|
|
ERVL |
0 |
0 bp |
0.00% |
|
|
ERVL |
2 |
118 bp |
0.00% |
|
|
ERVL-MaLRs |
0 |
0 bp |
0.00% |
|
|
ERVL-MaLRs |
0 |
0 bp |
0.00% |
|
|
ERV_classI |
0 |
0 bp |
0.00% |
|
|
ERV_classI |
0 |
0 bp |
0.00% |
|
|
ERV_classII |
0 |
0 bp |
0.00% |
|
|
ERV_classII |
0 |
0 bp |
0.00% |
|
DNAelements: |
10 |
4,735 bp |
2.59% |
|
DNAelements: |
39 |
3,894 bp |
0.03% |
||
|
|
hAT-Charlie |
4 |
2,073 bp |
1.13% |
|
|
hAT-Charlie |
27 |
2,315 bp |
0.02% |
|
|
TcMar-Tigger |
0 |
0 bp |
0.00% |
|
|
TcMar-Tigger |
1 |
61 bp |
0.00% |
|
Unclassified: |
0 |
0 bp |
0.00% |
|
Unclassified: |
0 |
0 bp |
0.00% |
||
|
Total interspersed repeats: |
|
5,594 bp |
3.06% |
|
Total interspersed repeats: |
|
6,363 bp |
0.05% |
||
|
Small RNA: |
4 |
257 bp |
0.14% |
|
Small RNA: |
6 |
2,083 bp |
0.01% |
||
|
Satellites: |
0 |
0 bp |
0.00% |
|
Satellites: |
0 |
0 bp |
0.00% |
||
|
Simple repeats: |
0 |
0 bp |
0.00% |
|
Simple repeats: |
1155 |
44,636 bp |
0.32% |
||
|
Low complexity: |
0 |
0 bp |
0.00% |
|
Low complexity: |
275 |
13,704 bp |
0.10% |
||
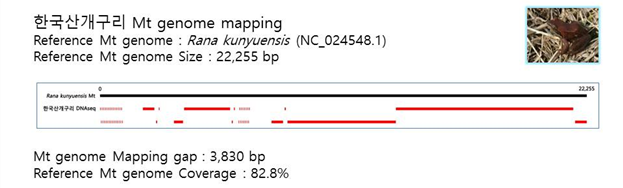
D. Mito-genome Map
(4) Microsatellite 후보군
A. SSR statistics
|
Motif(-mer) |
2 |
3 |
4 |
5 |
6 |
7 |
8 |
9 |
10 |
Total |
|
한국산개구리 |
69 |
11 |
8 |
- |
- |
- |
- |
- |
- |
88 |
B. 바코드서열
>한국산개구리(Rana coreana)
GGCATAATTGGAACAGCCCTAAGCCTCCTCATCCGAGCAGAACTCAGCCAGCCGGGGACC
CTCCTGGGGGATGATCAAATTTACAATGTCATCGTCACTGCCCATGCATTCGTAATAATT
TTTTTTATGGTTATACCAATTTTAATCGGGGGCTTTGGCAATTGGCTTGTCCCACTAATG
ATTGGAGCCCCCGACATAGCTTTCCCGCGAATAAACAACATAAGCTTCTGACTACTCCCA
CCTTCCTTTTTTCTTCTCTTGGCCTCTTCCACAGTTGAAGCAGGAGCAGGCACAGGCTGA
ACAGTCTACCCCCCTCTAGCTAGCAATCTAGCCCACGCTGGGCCATCAGTAGACATGGCC
ATCTTCTCTCTACACCTAGCCGGGGTATCATCCATTCTAGGGGCCATTAACTTTATCACA
ACAATTGTTAATATAAAACCCTCATCCACAACCCAGTACCAAACACCTCTCTTTGTATGA
TCCGTCTTAATTACTGCTGTTCTTCTACTTCTTTCCCTCCCCGTCCTGGCCGCCGGAATC
ACTATACTCCTCACAGACCGAAACCTAAACACTACCTTCTTCGATCCTGCCGGAGGCGGA