깊은산부전나비 (Protantigius superans)
|
|
▣ 국 명: 깊은산부전나비 ▣ 학 명: Protantigius superans ▣ 지정번호: 멸종위기동물 Ⅱ급 ▣ 계 통: 곤충강-나비목-부전나비과 |
◎ 생김새 : 날개를 편 길이가 37mm로 수컷에 비해 암컷이 조금 크다. 암컷 날개의 바깥 가장자리가 둥글며, 날개 윗면에 흰색 무늬가 있는 것이 특징이다.
◎ 서식지 : 높은 산지의 잡목림이나 그 주변 계곡에 서식하면서 높이 매달린 참나무의 잎 위에서 서식한다. 수컷은 오전에 높이가 낮은 풀 위에서 일광욕을 하며 한낮에 높은 나무위에서 머무는 것이 특징이다.
◎ 먹이 : 암컷의 경우 큰까치수염의 꽃에서 즙을 빨아먹는다.
◎ 번식 : 6~8월에 걸쳐 연 1회 산란한다.
◎ 분포 : 한국의 설악산?태백산?계룡산?소백산에서 분포가 확인되고 있으며 중국 서부, 아무르 지방 등에도 분포한다.
1) 채집정보
- 채집허가: 2014.06.30 / 5마리
- 채 집 자: 빛고을생태연구소 정헌천
2) 채집현황 및 샘플처리
|
깊은산부전나비 채집현황 |
샘플처리 |
|||
|
채집일 |
개체 수 |
DNAseq |
RNAseq |
표본 |
|
2014.07.30 |
5 EA |
3 EA |
2 EA |
- |
3) QC Data
|
샘플종 |
DNA QC |
RNA QC |
Result |
||
|
Concentration (ug/ul) |
Total Conc. (ug) |
Concentration (ug/ul) |
RIN (>7) |
||
|
깊은산부전나비 |
67 |
13.4 |
605 |
7.7 |
pass |
4) 기존 NCBI Taxonomy 유전정보 현황
|
유전정보 종류 |
Protantigius 속 유전자원 |
Protantigius superans 유전자원 |
|
Nucleotide |
2 |
2 |
|
Protein |
26 |
26 |
|
Genome |
1 |
1 |
|
Gene |
13 |
13 |
5) Sequencing 분석 결과
- Illumina DNA Sequncing 결과
|
Input information |
|||||
|
|
|
|
Region1 |
Region2 |
Total |
|
raw data |
|||||
|
Number Of Reads |
|
|
129,863,893 |
129,863,893 |
259,727,786 |
|
Number Of Bases |
|
|
16,362,850,518 |
16,362,850,518 |
32,725,701,036 |
|
error correction |
|||||
|
Number Of Reads |
|
|
129,112,091 |
128,020,730 |
257,132,821 |
|
Number Of Bases |
|
|
16,086,253,244 |
15,854,149,526 |
31,940,402,770 |
|
pair reads |
|||||
|
Number Of Reads |
|
|
|
|
254,559,484 |
|
Number Of Bases |
|
|
|
|
31,620,310,238 |
|
single reads |
|||||
|
Number Of Reads |
|
|
|
|
2,573,337 |
|
Number Of Bases |
|
|
|
|
320,092,532 |
|
Assembly Results |
|||||
|
Scaffold Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Scaffolds |
1,421,831 |
1,421,831 |
273,866 |
87,796 |
16,473 |
|
Number Of Bases |
497,650,177 |
497,650,177 |
271,752,744 |
142,465,164 |
46,860,340 |
|
Avg. Scaffold Size |
350 |
350 |
992 |
1,622 |
2,844 |
|
N50 Scaffold Size |
568 |
568 |
1,038 |
1,600 |
2,745 |
|
N80 Scaffold Size |
198 |
198 |
672 |
1,185 |
2,224 |
|
N90 Scaffold Size |
135 |
135 |
582 |
1,085 |
2,106 |
|
largest Scaffold Size |
25,015 |
25,015 |
25,015 |
25,015 |
25,015 |
|
Contig Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Contigs |
10,316,484 |
1,421,831 |
273,866 |
87,796 |
16,473 |
|
Number Of Bases |
883,912,259 |
497,650,177 |
271,752,744 |
142,465,164 |
46,860,340 |
|
Avg. Contig Size |
85 |
350 |
992 |
1,622 |
2,844 |
|
N50 Contig Size |
140 |
568 |
1,038 |
1,600 |
2,745 |
|
N80 Contig Size |
38 |
198 |
672 |
1,185 |
2,224 |
|
N90 Contig Size |
33 |
135 |
582 |
1,085 |
2,106 |
|
Largest Contig Size |
25,015 |
25,015 |
25,015 |
25,015 |
25,015 |
- Illumina RNA Sequncing 결과
|
Input information |
|||||
|
|
Region1 |
Region2 |
Region3 |
Region4 |
Total |
|
raw data |
|||||
|
Number Of Reads |
86,953,698 |
86,953,698 |
42,483,837 |
42,483,837 |
258,875,070 |
|
Number Of Bases |
10,956,165,948 |
10,956,165,948 |
5,352,963,462 |
5,352,963,462 |
32,618,258,820 |
|
error correction |
|||||
|
Number Of Reads |
86,247,434 |
85,658,620 |
42,041,849 |
41,735,911 |
255,683,814 |
|
Number Of Bases |
10,727,944,064 |
10,633,308,735 |
5,205,155,945 |
5,170,541,353 |
31,736,950,097 |
|
pair reads |
|||||
|
Number Of Reads |
|
170,754,850 |
|
83,210,134 |
253,964,984 |
|
Number Of Bases |
|
21,230,599,163 |
|
10,313,925,455 |
31,544,524,618 |
|
single reads |
|||||
|
Number Of Reads |
|
1,151,204 |
|
567,626 |
1,718,830 |
|
Number Of Bases |
|
130,653,636 |
|
61,771,843 |
192,425,479 |
|
Assembly Results |
|||||
|
Scaffold Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Scaffolds |
1,486,160 |
1,486,160 |
272,048 |
80,236 |
13,142 |
|
Number Of Bases |
500,048,358 |
500,048,358 |
258,627,728 |
125,944,334 |
36,571,418 |
|
Avg. Scaffold Size |
336 |
336 |
950 |
1,569 |
2,782 |
|
N50 Scaffold Size |
523 |
523 |
981 |
1,537 |
2,677 |
|
N80 Scaffold Size |
192 |
192 |
655 |
1,166 |
2,206 |
|
N90 Scaffold Size |
134 |
134 |
573 |
1,077 |
2,097 |
|
largest Scaffold Size |
25,015 |
25,015 |
25,015 |
25,015 |
25,015 |
|
Contig Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Contigs |
10,352,244 |
1,498,015 |
271,732 |
79,240 |
12,551 |
|
Number Of Bases |
886,544,103 |
500,253,443 |
256,378,111 |
123,288,925 |
34,571,699 |
|
Avg. Contig Size |
85 |
333 |
943 |
1,555 |
2,754 |
|
N50 Contig Size |
140 |
517 |
972 |
1,521 |
2,649 |
|
N80 Contig Size |
38 |
190 |
652 |
1,162 |
2,198 |
|
N90 Contig Size |
33 |
134 |
572 |
1,075 |
2,093 |
|
Largest Contig Size |
25,015 |
25,015 |
25,015 |
25,015 |
25,015 |
나) 멸종위기종 동물 Genome Survey
|
검수용 샘플 DNA 농도 및 총 양, 확증표본 현황표 |
|||||||
|
Sample명 |
농도(ng/ul) |
Volume(ul) |
DNA 총량(ug) |
O.D A260/280 |
확증표본 |
채집허가일 |
채집일 |
|
깊은산부전나비 |
751 |
996 |
747 |
2.02 |
○ |
2014.6.30 |
2014.7.30 |
(2) 유전체 서열확보 : 현재 20종 확보
|
Sample |
구분 |
raw data size |
trim data size |
비고 |
|
깊은산부전나비 |
DNA |
32,725,701,036 |
31,940,402,770 |
|
|
RNA |
32,618,258,820 |
31,736,950,097 |
|
(3) 기타분석
A. Genome Size estimation
|
샘 플 명 |
Kmer size |
Kmer total |
Pk depth |
Genome_size |
Coverage(X) |
|
깊은산부전나비 |
23 |
20,258,767,308 |
28 |
723,527,403 |
35.8974 |
B. 반복서열
- 시퀀싱이 완료된 깊은산부전나비 DNA 및 RNA의 contigs 중 1kb 이상의 contigs의 반복서열을 탐색(Left; Repeat masking results of DNA contigs, Right; Repeat masking results of RNA contigs).
|
sequences: 87,796 |
|
sequences: 6,332 |
||||||||
|
total length: 142,465,164 bp (142,465,164 bp excl N/X-runs) |
|
total length: 9,820,280 bp (9,820,280 bp excl N/X-runs) |
||||||||
|
GClevel:35.29% |
|
GC level: 39.00 % |
||||||||
|
bases masked: 54,158 bp ( 0.04 %) |
|
bases masked: 3,079 bp ( 0.03 %) |
||||||||
|
|
numberof elements |
length occupied |
percentage of sequence |
|
|
numberof elements |
length occupied |
percentage of sequence |
||
|
SINEs: |
19 |
941 bp |
0.00% |
|
SINEs: |
0 |
0 bp |
0.00% |
||
|
|
ALUs |
0 |
0 bp |
0.00% |
|
|
ALUs |
0 |
0 bp |
0.00% |
|
|
MIRs |
13 |
691 bp |
0.00% |
|
|
MIRs |
0 |
0 bp |
0.00% |
|
LINEs: |
139 |
12401 bp |
0.01% |
|
LINEs: |
9 |
601 bp |
0.01% |
||
|
|
LINE1 |
10 |
617 bp |
0.00% |
|
|
LINE1 |
2 |
116 bp |
0.00% |
|
|
LINE2 |
22 |
1442 bp |
0.00% |
|
|
LINE2 |
0 |
0 bp |
0.00% |
|
|
L3/CR1 |
65 |
4201 bp |
0.00% |
|
|
L3/CR1 |
6 |
355 bp |
0.00% |
|
LTRelements: |
12 |
757 bp |
0.00% |
|
LTRelements: |
1 |
45 bp |
0.00% |
||
|
|
ERVL |
2 |
71 bp |
0.00% |
|
|
ERVL |
0 |
0 bp |
0.00% |
|
|
ERVL-MaLRs |
0 |
0 bp |
0.00% |
|
|
ERVL-MaLRs |
0 |
0 bp |
0.00% |
|
|
ERV_classI |
6 |
394 bp |
0.00% |
|
|
ERV_classI |
1 |
45 bp |
0.00% |
|
|
ERV_classII |
0 |
0 bp |
0.00% |
|
|
ERV_classII |
0 |
0 bp |
0.00% |
|
DNAelements: |
201 |
31416 bp |
0.02% |
|
DNAelements: |
16 |
2210 bp |
0.02% |
||
|
|
hAT-Charlie |
62 |
9596 bp |
0.01% |
|
|
hAT-Charlie |
4 |
307 bp |
0.00% |
|
|
TcMar-Tigger |
18 |
2500 bp |
0.00% |
|
|
TcMar-Tigger |
2 |
225 bp |
0.00% |
|
Unclassified: |
39 |
2830 bp |
0.00% |
|
Unclassified: |
3 |
223 bp |
0.00% |
||
|
Total interspersed repeats: |
|
48345 bp |
0.03% |
|
Total interspersed repeats: |
|
3079 bp |
0.03% |
||
|
Small RNA: |
104 |
5884 bp |
0.00% |
|
Small RNA: |
0 |
0 bp |
0.00% |
||
|
Satellites: |
0 |
0 bp |
0.00% |
|
Satellites: |
0 |
0 bp |
0.00% |
||
|
Simple repeats: |
0 |
0 bp |
0.00% |
|
Simple repeats: |
0 |
0 bp |
0.00% |
||
|
Low complexity: |
0 |
0 bp |
0.00% |
|
Low complexity: |
0 |
0 bp |
0.00% |
||
D. Mito-genome Map
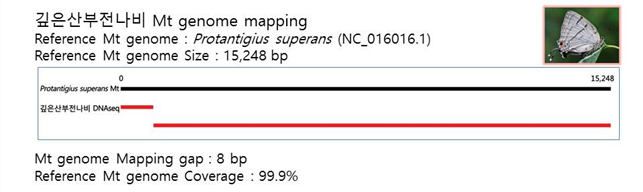
(4) Microsatellite 후보군
A. SSR statistics
|
Motif(-mer) |
2 |
3 |
4 |
5 |
6 |
7 |
8 |
9 |
10 |
Total |
|
깊은산부전나비 |
15,573 |
9,416 |
536 |
72 |
24 |
2 |
- |
- |
- |
25,623 |
B. 바코드서열
>깊은산부전나비(Protantigius superans)
AAAAAAAATATTACGCTGTTATCCCTAAGGTAATTTTTTCTTTTAATCATTAATTATGGA
TCATTTATTCATTTATTTTTGTTTAATTTTAAAAAAAGTTTATTTAATTTTTTTATCACC
CCAATACAATAATTATTTATCTTAAATATTAAATTTTTATATAAATATTTTAAATTAATA
ATTATAAAACTCTATAGGGTCTTCTCGTCTTTTAATTTTATTTTAGCTTTTTAACTAAAA
AATTAAATTCTATAAATAACTAAGAGACAGTTTATATTTCATCAAATCTTTCATACAAGT
CTTCAATTAAAAGACTAATGATTATGCTACCTTTGTACAGTCAATATACTGCAGCCCTTT
AATAATTAAATCAGTGGGCAGATTAGACTTTAAATTATTTTTCAAAAAGACATGTTTTTG
TTAAACAGGTGAATATATTAATTTGCCGAATTCCTTTTAATTATATTTTAATAATTCATA
TTTATTTTAAATTATTTTATTATACTAATTTTATCATTTTTTTTAATATTATTTAATTAA
AAATTATATTTTTATAAAAAATTAAATTCATATTAAATTTTATTCTAAATAAAATTTTTT
ATAATAAACTTTAATTTTTAAACAATATAAATTTTAAAAATTTTAT