수염풍뎅이 Polyphylla laticollis manchurica (Semenov,1900)
|
|
▣ 국 명 : 수염풍뎅이 ▣ 학 명 : Polyphylla laticollis manchurica Semenov,1900 ▣ 지정번호 : 멸종위기 야생생물 Ⅰ급 ▣ 계 통 : 절지동물문 > 곤충강 > 딱정벌레목 > 검정풍뎅이과 > 수염풍뎅이속 |
◎ 형 태 : 크기는 약 33~37mm 정도의 크기로 몸의 색은 진한 갈색으로 불규칙한 흰색의 얼룩무늬가 곳곳에 나타나 있으며, 머리는 작고 머리방패는 긴사각형으로 앞가두리는 넓고 위로 젖혀져 있으며, 회백색과 황백색의 짧은 털이 촘촘히 나타나 있다. 촉각은 10마디로 수컷의 곤봉부는 7마디로서 매우 길고 굽어 있으며 암컷은 5~6마디로 직선형이다. 앞가슴 등판은 짧고 넓으며 흰색의 짧은 털이 있고 뒷가두리의 아래쪽에 연한 노란색의 짧은 털이 촘촘히 있다. 딱지날개는 타원형으로 분명하지 않은 3줄의 세로로 솟아오른 부분을 찾아볼 수 있다. 수컷의 앞다리 종아리마디에는 2개의 가시돌기가 있고, 암컷에는 3개의 가시돌기가 있다.

◎ 생 태 : 평지의 경작지 주변의 풀밭에서 살아가고 있으며 늦은 봄부터 가을까지 찾아볼 수 있다. 특히 6~7월에 개체수가 많아지고, 주로 파꽃에 무리를 지어 모이기도 하며 불빛에 날아든다. 유충은 갈대 따위의 하천변 식물이 퇴적한 곳에 사는 것으로 알려져 있으며, 소나무류, 사시나무류, 갈참나무, 참나무 등의 뿌리를 갉아먹는다. 강변이나 하천가의 환경 개조로 인하여 서식처가 없어져 개체수가 급격히 감소하고 있다.
◎ 분 포 : 우리나라와 일본, 중국 동북과 몽골에 분포한다.
① 채집정보
- 채 집 자 : 빛고을생태연구소 정헌천
- 채집지역 : 충청남도 논산시
|
|
|
|
수염풍뎅이 서식지 및 포획장소 환경 |
채집한 수염풍뎅이 1개체 |
② 채집현황 및 샘플처리
|
수염풍뎅이 채집현황 |
샘플처리 |
|||
|
채집일 |
개체 수 |
DNAseq |
RNAseq |
표본 |
|
2017.07.14 |
1 EA |
1 EA |
ㅡ |
|
③ NCBI Taxonomy 유전정보 현황
|
유전정보 종류 |
Polyphylla 속 유전자원 |
Polyphylla laticollis manchurica 유전자원 |
|
Nucleotide |
15 |
1 |
|
Protein |
21 |
13 |
④ Sequencing 결과
|
Type |
Total Reads |
Total Bases |
Total Bases (Gb) |
GC Rate |
비고 |
|
DNA |
255,400,882 |
38,565,533,182 |
38.5 |
34.6 |
|
|
RNA |
53,433,048 |
7,587,425,115 |
7.5 |
37.3 |
|
- Illumina DNA Sequencing 결과
|
SOAP de novo |
|||||
|
Pl_DNA hiseq |
|||||
|
Input information |
|||||
|
|
|
|
Region1 |
Region2 |
Total |
|
raw data |
|||||
|
Number Of Reads |
|
|
127,700,441 |
127,700,441 |
255,400,882 |
|
Number Of Bases |
|
|
19,282,766,591 |
19,282,766,591 |
38,565,533,182 |
|
error correction |
|||||
|
Number Of Reads |
|
|
127,117,664 |
124,428,336 |
251,546,000 |
|
Number Of Bases |
|
|
18,583,153,254 |
17,624,105,232 |
36,207,258,486 |
|
pair reads |
|||||
|
Number Of Reads |
|
|
|
248,324,094 |
248,324,094 |
|
Number Of Bases |
|
|
|
35,776,477,269 |
35,776,477,269 |
|
single reads |
|||||
|
Number Of Reads |
|
|
|
3,221,906 |
3,221,906 |
|
Number Of Bases |
|
|
|
430,781,217 |
430,781,217 |
|
Assembly Results |
|||||
|
Scaffold Metrics |
> 100 bp |
> 500 bp |
> 1000 bp |
> 2000 bp |
|
|
Number Of Scaffold |
|
1,175,590 |
388,863 |
227,233 |
92,844 |
|
Number Of Bases |
|
775,257,354 |
618,997,508 |
503,729,116 |
313,503,315 |
|
Avg. Scaffold Size |
|
659 |
1,591 |
2,216 |
3,376 |
|
N50 Scaffold Size |
|
1,567 |
2,027 |
2,445 |
3,411 |
|
N80 Scaffold Size |
|
495 |
1,039 |
1,487 |
2,454 |
|
N90 Scaffold Size |
|
245 |
758 |
1,236 |
2,213 |
|
Largest Scaffold Size |
|
27,462 |
27,462 |
27,462 |
27,462 |
|
Contigs |
> 0 bp |
> 100 bp |
> 500 bp |
> 1000 bp |
> 2000 bp |
|
Number Of Contigs |
29,305,583 |
2,374,859 |
303,967 |
112,520 |
18,070 |
|
Number Of Bases |
1,869,069,590 |
681,877,375 |
309,391,201 |
174,952,156 |
48,034,316 |
|
Avg. Contig Size |
63 |
287 |
1,017 |
1,554 |
2,658 |
|
N50 Contig Size |
66 |
418 |
1,098 |
1,533 |
2,556 |
|
N80 Contig Size |
36 |
153 |
702 |
1,175 |
2,174 |
|
N90 Contig Size |
34 |
120 |
594 |
1,084 |
2,081 |
|
largest Contig Size |
11,939 |
11,939 |
11,939 |
11,939 |
11,939 |
- Illumina RNA Sequencing 결과
|
Total number of raw reads |
|
|
- Number of sequences |
53,433,048 |
|
- Number of bases |
7,587,425,115 |
|
Contig information |
|
|
- Total number of contig |
130,004 |
|
- Number of bases |
86,251,144 |
|
- Mean length of contig (bp) |
663.4 |
|
- N50 length of contig (bp) |
926 |
|
- GC % of contig |
35.20 |
|
- Largest contig (bp) |
14,041 |
|
- No. of large contigs (≥500bp) |
51,955 |
|
Transdecoder information |
|
|
- Total number of contig |
75,219 |
|
- Number of bases |
76,894,513 |
|
- Mean length of contig (bp) |
1,022.3 |
|
- N50 length of contig (bp) |
1,537 |
|
- GC % of contig |
36.04 |
|
- Largest contig (bp) |
14,041 |
|
- No. of large contigs (≥500bp) |
46,462 |
|
Unigene information |
|
|
- Total number of unigenes |
18,172 |
|
- Number of bases |
25,001,252 |
|
- Mean length of unigene (bp) |
1,375.8 |
|
- N50 length of unigene (bp) |
1,815 |
|
- GC % of unigene |
35.89 |
|
- Length ranges (bp) |
224 – 17,572 |
2) 현재 상황에 대한 고찰
가) 대상종 및 시료확보 상황
(1) 초기 예비종을 포함한 채집 대상종 : 23종(절지동물(곤충) 9종, 연체동물 8종, 양서류 6종)
(2) 채집 완료 종 : 9종(절지동물(곤충) 5종, 연체동물 1종, 양서류 3종)
나) 멸종위기동물의 Genome survey
(1) Library QC data
- DNA Library QC
|
Delivery name |
Tape Concentration (ng/ul) |
Volume (ul) |
Tape Quantity (ug) |
Main peak Size (bp) |
Result |
|
수염풍뎅이 |
87.237 |
80 |
6.979 |
470 |
pass |
|
|
- RNA Library QC
|
Delivery name |
Tape Concentration (ng/ul) |
Volume (ul) |
Tape Quantity (ug) |
Main peak Size (bp) |
Result |
|
수염풍뎅이 |
325.732 |
30 |
9.772 |
288 |
pass |
|
|
(2) 고품질 genomic DNA 채취 및 표본확보
- 검수용 DNA는 대부분 만족하였고, 멸종위기종인 대상종들은 채집량의 제한으로 인해 확증표본으로 제출 가능한 표본을 확보하지 못하여 사진으로 대체하고자 함.
|
검수용 샘플 DNA 농도 및 총 양 현황표 |
||||||
|
Sample 명 |
농도(ng/ul) |
Volume(ul) |
DNA 총량(ug) |
O.D A260/280 |
추출일자 |
저장장소 |
|
수염풍뎅이 |
589 |
300 |
176.7 |
1.94 |
170927 |
1XTE pH8.0 |
(3) 기타분석
A. Genome Size estimation – Jellyfish 프로그램
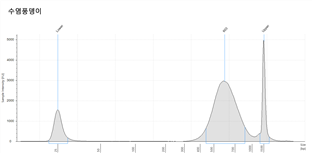
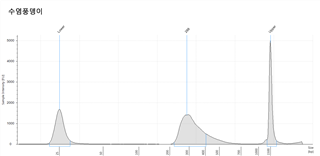
각 대상종을 대상으로 Jellyfish 프로그램으로 Genome size를 예측한 결과는 아래와 같다. 각 분석은 k-mer 값 17mer, 19mer, 21mer를 대상으로 하였고 그것을 통해 예측된 genome의 크기는 Estimation genome size 에 표시되었고, 이를 통해 이번 사업에서 분석한 서열의 양의 coverage는 Genome coverage depth(Coverage Depth)에 표기되었다. 그리고 각 종의 표 아래 그래프는 kmer값 별로 나타난 depth의 예측 그래프로 peak값을 확인할 수 있다.
|
샘플명 |
K-mer (K-mersize) |
Estimateddepth (K-merCoveragedepth) |
Genomecoveragedepth (Coveragedepth) |
Estimatedgenome size |
|
수염풍뎅이 |
17 |
23 |
25.73 |
749,546,066.7 |
|
19 |
22 |
24.98 |
772,007,211.5 |
|
|
21 |
22 |
25.36 |
760,398,080.5 |
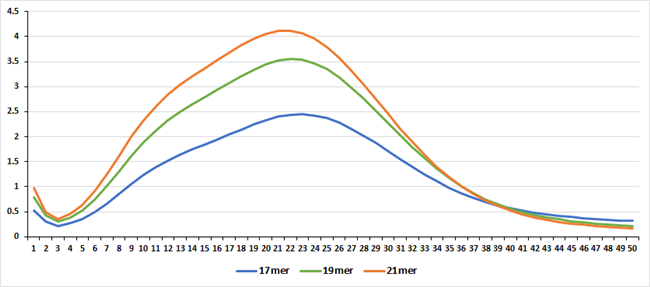
B. 반복서열
- 시퀀싱이 완료된 샘플 DNA 의 contigs 중 1kb 이상의 contigs의 반복서열을 탐색 (Repeat masking results of DNA contigs)
|
수염풍뎅이 (Polyphylla laticollis manchurica) |
|
수염풍뎅이 (Polyphylla laticollis manchurica) 는 1kb 이상의 contig 의 반복서열은 18,172개 read (25,001,252 bp) 이고, GC level 은 35.89%이다. Repeat Masking 으로 나온 반복서열은 341,632 bp (전체의 1.37%) 만큼 있다. Small RNA 은 33개 read (10,135 bp) 가 있고, Simple repeat 는 4,140개 read (185,493 bp) 가 있다. |
||||
|
sequences: 18,172 |
|
|||||
|
total length: 25,001,252 bp (25,001,252 bp excl N/X-runs) |
|
|||||
|
GC level: 35.89 % |
|
|||||
|
bases masked: 341,632 bp ( 1.37%) |
|
|||||
|
|
number of elements |
length occupied |
percentage of sequence |
|
||
|
|
||||||
|
SINEs: |
3 |
191 bp |
0.00% |
|
||
|
|
ALUs |
0 |
0 bp |
0.00% |
|
|
|
|
MIRs |
2 |
132 bp |
0.00% |
|
|
|
LINEs: |
55 |
4,849 bp |
0.02% |
|
||
|
|
LINE1 |
8 |
506 bp |
0.00% |
|
|
|
|
LINE2 |
5 |
242 bp |
0.00% |
|
|
|
|
L3/CR1 |
27 |
1,727 bp |
0.01% |
|
|
|
LTRelements: |
9 |
1,238 bp |
0.00% |
|
||
|
|
ERVL |
2 |
80 bp |
0.00% |
|
|
|
|
ERVL-MaLRs |
0 |
0 bp |
0.00% |
|
|
|
|
ERV_classI |
1 |
53 bp |
0.00% |
|
|
|
|
ERV_classII |
0 |
0 bp |
0.00% |
|
|
|
DNAelements: |
403 |
81,584 bp |
0.33% |
|
||
|
|
hAT-Charlie |
59 |
14,668 bp |
0.06% |
|
|
|
|
TcMar-Tigger |
104 |
30,119 bp |
0.12% |
|
|
|
Unclassified: |
6 |
581 bp |
0.00% |
|
||
|
Total interspersed repeats: |
|
88,443 bp |
0.35% |
|
||
|
Small RNA: |
33 |
10,135 bp |
0.04% |
|
||
|
Satellites: |
1 |
45 bp |
0.00% |
|
||
|
Simple repeats: |
4,140 |
185,493 bp |
0.74% |
|
||
C. Mito-genome Map
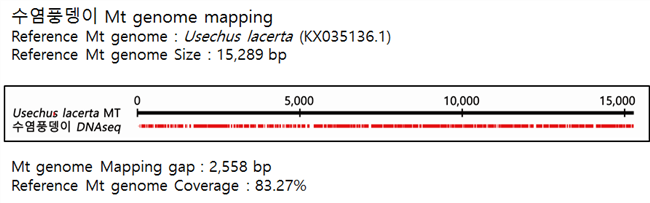
수염풍뎅이는 Reference로 15,289 bp 크기의 Usechus lacerta 미토콘드리아 genome을 이용하여 맵핑하였다. 갭은 2,558 bp 로, Coverage 는 83.27%로 확인되었다.
D. Transcriptome data 분석
- 수염풍뎅이
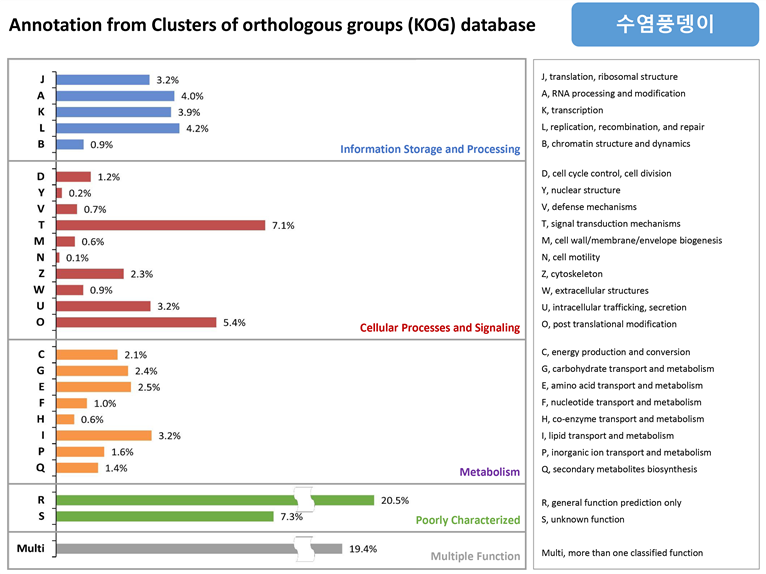
수염풍뎅이의 KOG 결과, “Infomation Storage and Processing” 부분에서는 replication, recombination and repair 관련이 4.2%로 가장 많이 매칭되고, “Cellular Processes and Signaling” 에서는 signal transduction mechanisms (7.1%), post translational modification (5.4%) 로 많이 매칭된다. “Metabolism” 부분은 평균 1% 대로 낮게 매칭되고, Multiple Function 은 19.4% 이다.
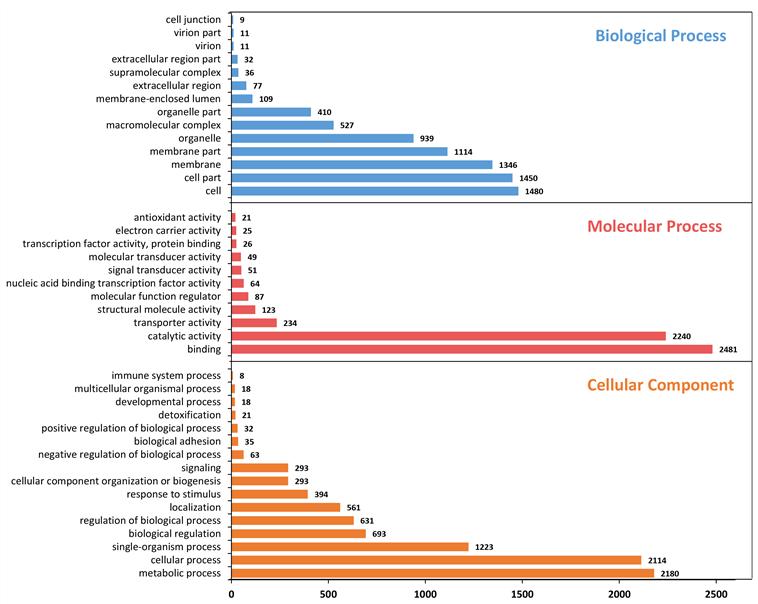
수염풍뎅이의 GO 결과, Biological Process에서 cell (1,480개), cell part (1,450개) 순으로 많이 매칭된다. Molecular Function 에서는 binding (2,481개), catalytic activity (2,240개) 순으로 가장 많이 매칭된다. 또한, Cellular Component 부분에서는 metabolic process (2,180개), cellular process (2,114개), single-organism process (1,223개) 순으로 많이 매칭된다.
(4) Microsatellite 후보군
A. SSR statistics
|
수염풍뎅이 |
||||||||||||
|
Repeats |
4 |
5 |
6 |
7 |
8 |
9 |
10 |
11 |
12 |
13 |
… |
Total |
|
Di |
0 |
0 |
178 |
75 |
77 |
34 |
31 |
12 |
16 |
4 |
|
471 |
|
Tri |
0 |
203 |
69 |
13 |
9 |
5 |
2 |
2 |
0 |
0 |
|
303 |
|
Tetra |
55 |
14 |
4 |
0 |
0 |
1 |
0 |
0 |
0 |
1 |
|
75 |
|
Penta |
2 |
1 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
|
3 |
|
Hexa |
2 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
|
2 |
|
Total |
59 |
218 |
251 |
88 |
86 |
40 |
33 |
14 |
16 |
5 |
|
854 |
→ SSR 검색결과 총 854개의 SSR 서열을 확인할 수 있었으며, Di motif 가 471개로 가장 많이 존재하는 것을 확인 할 수 있었으며, 이를 기반으로 하여 SSR marker 후보군에 대한 primer 를 2,326개를 디자인 하였다.
B. Primer sequence
→ 각 종의 유전자를 분석하여 그 종을 구별할 수 있는 primer를 만든다. primer 의 조건은 Motif 약 4~6개이며, Forward 와 Reverse primer 가 18~22개 정도이고 Tm값이 54.5~55.5 정도가 적당하다.
- 수염풍뎅이
|
Sequence |
Motif |
Forward Primer |
Tm |
|
Reverse Primer |
|||
|
Pl_Uni_00241 |
(AT)27 |
ACGAACATGGAAATTCACTT |
54.61 |
|
GTAGCTCAGGAGTGGGTCATA |
56.87 |
||
|
Pl_Uni_00452 |
(AC)10 |
AACACACACACACAAACACAC |
55.12 |
|
AACCTGTATCCAAAGGAAATC |
54.84 |
||
|
Pl_Uni_00595 |
(ATA)7 |
GCTGCAGTGTTTAGGAACC |
55.9 |
|
TGAAACAATTGAACGTCTTCT |
55.02 |
||
|
Pl_Uni_00701 |
(TTA)6 |
GGTTCATAAACACGATGCTAA |
55.43 |
|
AGTCGAACCAAATAACGATAG |
53.75 |
||
|
Pl_Uni_00703 |
(AAT)6 |
CATATCCAGGTTCATTAGCAG |
54.96 |
|
CCCTTGAATTCTTTAGGATGT |
55.01 |
||
|
Pl_Uni_00815 |
(ACAAAA)3 |
ACATGGAAATACCAGAAGGTT |
55.18 |
|
CAACCCAATTTCTTTCAGATA |
54.5 |
||
|
Pl_Uni_00899 |
(AC)11 |
GACAATCACACAAAAACACAA |
54.48 |
|
GTTGTTTGAGTCAAGCCTTAG |
54.37 |
||
|
Pl_Uni_00910 |
(AT)15 |
TCCTGAACAAAACTGAGGTTA |
55.03 |
|
AGACAACAACGGTTCTCTTG |
55.36 |
||
|
Pl_Uni_00949 |
(TA)22 |
CCAATGGATGGACTGTTATAC |
54.45 |
|
CCAATGGATGGACTGTTATAC |
54.45 |
||
|
Pl_Uni_00992 |
(AT)11 |
GTATTTCCACATTCCAAAACA |
55.08 |
|
CAATAGCGAAAACTAGCAGAA |
55.24 |
||
|
Pl_Uni_04064 |
(TAA)6 |
AATATATACGCGCATTTTCA |
53.71 |
|
AGCACATTTTTACGATCTGTT |
54.19 |
||
|
Pl_Uni_04103 |
(TCT)6 |
GTTACTTGTGCATAGGGCATA |
55.47 |
|
TGCGAAAATAATGAAGACCTA |
55.25 |
||
|
Pl_Uni_04112 |
(AT)9 |
ATATCCCCATAAGAAGTGGTC |
54.64 |
|
GATGTGATTGTACGTTGAGGT |
54.99 |
||
|
Pl_Uni_04153 |
(CAA)6 |
AAAAGACGTCCCTAAATTCAA |
55.63 |
|
GTTTTAGCGTTGTTCAAACTG |
55.31 |
||
|
Pl_Uni_04438 |
(TAA)6 |
AGTGTTTCGTGGTGTACTGAG |
55.32 |
|
CACTGTGCACGTTACAATTAG |
54.51 |
||
|
Pl_Uni_04446 |
(ACA)6 |
CAATCACAACAACAGCAGAC |
55.08 |
|
GCCTTGTATAGAGTGCATAGC |
54.46 |
C. 바코드 서열
올해 17년도에 채집한 6종의 유전자를 분석하여 그 종을 구별할 수 있는 Barcode 서열을 만들었고, 그 Barcode 서열을 가지고 NCBI (National Center for Biotechnology Information) 사이트에 있는 BLAST (Basic Local Alignment Search Tool)을 이용하여 결과를 확인하였다. 각 종의 서열들의 BLAST 결과 마커로서 주로 사용되는 COI 서열과 매치되는 것으로 보아 바코드 서열로 사용가능할 것으로 사료된다.
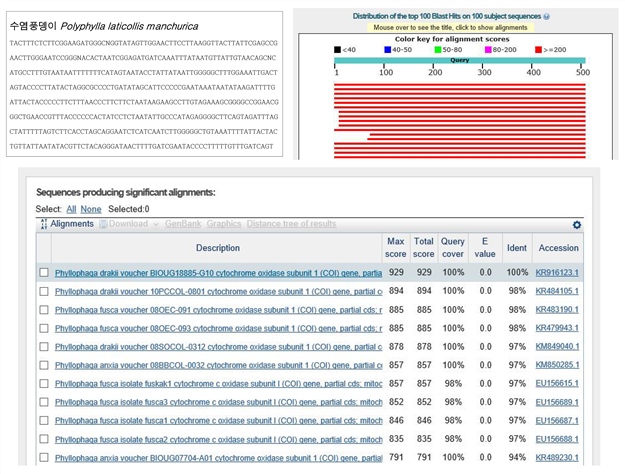
(5) Gene prediction
|
샘플종 |
Number of Contig (>1kb) |
Number of Bases (>1kb) |
N50 Contig Size |
Number of gene prediction |
|
수염풍뎅이 |
9,854 |
20,018,570 |
1,815 |
9,061 |
각종별로 transcriptome 서열을 대상으로 1kb 이상만을 선택하여 Gene predcition 한 결과 위의 매칭된 contig, base 수는 위의 표와 같고 N50되는 Contig의 길이도 보통 1~3kb 정도로 나타났다. 그리고 이들을 통해 예측된 유전자의 수는 8천개에서 4만개 이상으로 다양하게 예측되었다.
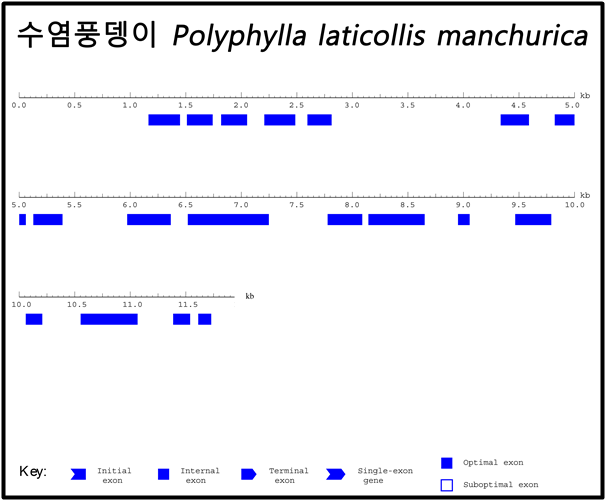
※ Long contig의 Gene prediction 결과 그림 예시