금개구리 (Pelophylax chosenicus)
|
|
▣ 국 명: 금개구리 ▣ 영 명: Korean Golden frog ▣ 학 명: Pelophylax chosenicus ▣ 지정번호: 멸종위기야생동물 Ⅱ급 ▣ 계 통: 양서강-무미목-개구리과 |
◎ 생김새 : 등은 전반적으로 밝은 녹색, 배는 노란색을 띤다. 체형은 유선형이고 크기는 4~6cm 정도이다. 몸의 크기는 참개구리와 거의 같고 그 모양 또한 유사하나 등 옆선을 이루는 두 줄의 융기가 금색으로 돌출되어 있어 구별이 가능하다. 암?수 모두 울음주머니가 없는 것이 특징이다.
◎ 서식지 : 주로 서해안을 따라 저습지와 논 등지에서 서식하며 산란장소와 서식지를 공유한다. 주로 물에서 생활하며 위험을 느낄 때는 물속으로 뛰어들어 진흙에 몸을 감추기도 한다.
◎ 먹이 : 주로 곤충과 같은 절지동물을 섭식하며 거미류도 포식한다.
◎ 번식 : 산란 시기는 지역에 따라 차이가 있으며 대개 5월에서 7월에 번식한다.
◎ 분포 : 우리나라의 전역에 분포하였으나 서식지 파괴 및 농약살포로 개체수가 현저히 감소하였다. 현재 인천, 경기도, 충남, 경남 합천 지역에서 서식이 확인되고 있다.
1) 채집정보
- 채집허가: 2014.07.17 / 3마리
- 채 집 자: 서울대학교 민미숙
2) 채집현황 및 샘플처리
|
금개구리 채집현황 |
샘플처리 |
|||
|
채집일 |
개체 수 |
DNAseq |
RNAseq |
표본 |
|
2014.07.30 |
3 EA |
2 EA (몸통) |
2 EA (내장) |
1 EA, 70% EtOH |
3) QC Data
|
샘플종 |
DNA QC |
RNA QC |
Result |
||
|
Concentration (ug/ul) |
Total Conc. (ug) |
Concentration (ug/ul) |
RIN (>7) |
||
|
금개구리 |
168.7 |
30.3 |
588.2 |
8 |
pass |
4) 기존 NCBI Taxonomy 유전정보 현황
|
유전정보 종류 |
Pelophylax 속 유전자원 |
Pelophylax chosenicus 유전자원 |
|
Nucleotide |
6,006 |
292 |
|
Protein |
5,268 |
125 |
|
Genome |
13 |
- |
|
Gene |
421 |
12 |
5) Sequencing 분석 결과
- Illumina DNA Sequncing 결과
|
Input information |
|||||
|
|
|
|
Region1 |
Region2 |
Total |
|
raw data |
|||||
|
Number Of Reads |
|
|
191,514,083 |
191,514,083 |
383,028,166 |
|
Number Of Bases |
|
|
24,130,774,458 |
24,130,774,458 |
48,261,548,916 |
|
error correction |
|||||
|
Number Of Reads |
|
|
180,760,039 |
178,205,456 |
358,965,495 |
|
Number Of Bases |
|
|
20,804,185,966 |
20,391,011,834 |
41,195,197,800 |
|
pair reads |
|||||
|
Number Of Reads |
|
|
|
|
341,805,376 |
|
Number Of Bases |
|
|
|
|
39,508,954,354 |
|
single reads |
|||||
|
Number Of Reads |
|
|
|
|
17,160,119 |
|
Number Of Bases |
|
|
|
|
1,686,243,446 |
|
Assembly Results |
|||||
|
Scaffold Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Scaffolds |
3,917,707 |
3,917,707 |
625,561 |
111,120 |
5,586 |
|
Number Of Bases |
1,193,664,158 |
1,193,664,158 |
494,890,520 |
147,341,835 |
13,292,621 |
|
Avg. Scaffold Size |
304 |
304 |
791 |
1,325 |
2,379 |
|
N50 Scaffold Size |
418 |
418 |
788 |
1,283 |
2,304 |
|
N80 Scaffold Size |
193 |
193 |
594 |
1,091 |
2,093 |
|
N90 Scaffold Size |
131 |
131 |
544 |
1,042 |
2,045 |
|
largest Scaffold Size |
12,888 |
12,888 |
12,888 |
12,888 |
12,888 |
|
Contig Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Contigs |
74,088,165 |
6,311,447 |
211,521 |
9,614 |
72 |
|
Number Of Bases |
3,971,713,837 |
1,218,958,721 |
139,064,558 |
11,397,951 |
180,329 |
|
Avg. Contig Size |
53 |
193 |
657 |
1,185 |
2,504 |
|
N50 Contig Size |
50 |
204 |
637 |
1,144 |
2,378 |
|
N80 Contig Size |
33 |
127 |
544 |
1,046 |
2,101 |
|
N90 Contig Size |
32 |
111 |
520 |
1,021 |
2,044 |
|
Largest Contig Size |
5,517 |
5,517 |
5,517 |
5,517 |
5,517 |
- Illumina RNA Sequncing 결과
|
Input information |
|||||
|
|
|
|
Region1 |
Region2 |
Total |
|
raw data |
|||||
|
Number Of Reads |
|
|
140,093,046 |
140,093,046 |
280,186,092 |
|
Number Of Bases |
|
|
17,651,723,796 |
17,651,723,796 |
35,303,447,592 |
|
error correction |
|||||
|
Number Of Reads |
|
|
138,052,733 |
136,753,947 |
274,806,680 |
|
Number Of Bases |
|
|
17,111,028,619 |
16,917,671,690 |
34,028,700,309 |
|
pair reads |
|||||
|
Number Of Reads |
|
|
|
|
272,287,600 |
|
Number Of Bases |
|
|
|
|
33,752,871,362 |
|
single reads |
|||||
|
Number Of Reads |
|
|
|
|
2,519,080 |
|
Number Of Bases |
|
|
|
|
275,828,947 |
|
Assembly Results |
|||||
|
Scaffold Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Scaffolds |
169,398 |
169,398 |
34,901 |
15,461 |
5,195 |
|
Number Of Bases |
72,719,541 |
72,719,541 |
45,075,088 |
31,605,804 |
17,388,315 |
|
Avg. Scaffold Size |
429 |
429 |
1,291 |
2,044 |
3,347 |
|
N50 Scaffold Size |
785 |
785 |
1,553 |
2,177 |
3,395 |
|
N80 Scaffold Size |
244 |
244 |
797 |
1,361 |
2,421 |
|
N90 Scaffold Size |
158 |
158 |
630 |
1,164 |
2,198 |
|
largest Scaffold Size |
17,029 |
17,029 |
17,029 |
17,029 |
17,029 |
|
Contig Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Contigs |
2,700,519 |
240,510 |
32,133 |
10,912 |
2,318 |
|
Number Of Bases |
181,187,643 |
73,460,317 |
32,925,943 |
18,341,430 |
6,697,539 |
|
Avg. Contig Size |
67 |
305 |
1,024 |
1,680 |
2,889 |
|
N50 Contig Size |
63 |
424 |
1,098 |
1,681 |
2,802 |
|
N80 Contig Size |
36 |
171 |
675 |
1,210 |
2,252 |
|
N90 Contig Size |
32 |
130 |
580 |
1,095 |
2,119 |
|
Largest Contig Size |
9,924 |
9,924 |
9,924 |
9,924 |
9,924 |
|
Sample |
구분 |
raw data size |
trim data size |
비고 |
|
금개구리 |
DNA |
48,261,548,916 |
41,195,197,800 |
|
|
RNA |
35,303,447,592 |
34,028,700,309 |
|
(3) 기타분석
A. Genome Size estimation
|
샘 플 명 |
Kmer size |
Kmer total |
Pk depth |
Genome_size |
Coverage(X) |
|
금개구리 |
23 |
29,876,196,948 |
3 |
9,958,732,316 |
3.84615 |
B. 반복서열
- 시퀀싱이 완료된 금개구리 DNA 및 RNA의 contigs 중 1kb 이상의 contigs의 반복서열을 탐색 (Left; Repeat masking results of DNA contigs, Right; Repeat masking results of RNA contigs).
|
sequences: 9,614 |
|
sequences: 10,912 |
||||||||
|
total length: 11,397,951 bp (11,397,951 bp excl N/X-runs) |
|
total length: 18,341,430 bp (18,341,430 bp excl N/X-runs) |
||||||||
|
GClevel: 42.49% |
|
GC level: 44.90% |
||||||||
|
bases masked: 27,311 bp ( 0.24%) |
|
bases masked: 19,949 bp ( 0.11%) |
||||||||
|
|
numberof elements |
length occupied |
percentage of sequence |
|
|
numberof elements |
length occupied |
percentage of sequence |
||
|
SINEs: |
8 |
733 bp |
0.01% |
|
SINEs: |
3 |
285 bp |
0.00% |
||
|
|
ALUs |
0 |
0 bp |
0.00% |
|
|
ALUs |
0 |
0 bp |
0.00% |
|
|
MIRs |
3 |
166 bp |
0.00% |
|
|
MIRs |
0 |
0 bp |
0.00% |
|
LINEs: |
81 |
9,225 bp |
0.08% |
|
LINEs: |
46 |
5,211 bp |
0.03% |
||
|
|
LINE1 |
42 |
4,428 bp |
0.04% |
|
|
LINE1 |
20 |
2,311 bp |
0.01% |
|
|
LINE2 |
20 |
2,836 bp |
0.02% |
|
|
LINE2 |
10 |
919 bp |
0.01% |
|
|
L3/CR1 |
14 |
1,620 bp |
0.01% |
|
|
L3/CR1 |
13 |
1,822 bp |
0.01% |
|
LTRelements: |
24 |
6,112 bp |
0.05% |
|
LTRelements: |
9 |
1,887 bp |
0.01% |
||
|
|
ERVL |
0 |
0 bp |
0.00% |
|
|
ERVL |
1 |
40 bp |
0.00% |
|
|
ERVL-MaLRs |
0 |
0 bp |
0.00% |
|
|
ERVL-MaLRs |
0 |
0 bp |
0.00% |
|
|
ERV_classI |
3 |
270 bp |
0.00% |
|
|
ERV_classI |
4 |
1,171 bp |
0.01% |
|
|
ERV_classII |
0 |
0 bp |
0.00% |
|
|
ERV_classII |
1 |
83 bp |
0.00% |
|
DNAelements: |
88 |
8,417 bp |
0.07% |
|
DNAelements: |
100 |
11,350 bp |
0.06% |
||
|
|
hAT-Charlie |
47 |
3,838 bp |
0.03% |
|
|
hAT-Charlie |
74 |
6,662 bp |
0.04% |
|
|
TcMar-Tigger |
24 |
2,877 bp |
0.03% |
|
|
TcMar-Tigger |
4 |
509 bp |
0.00% |
|
Unclassified: |
1 |
76 bp |
0.00% |
|
Unclassified: |
1 |
67 bp |
0.00% |
||
|
Total interspersed repeats: |
|
24,563 bp |
0.22% |
|
Total interspersed repeats: |
|
18,800 bp |
0.10% |
||
|
Small RNA: |
22 |
3,210 bp |
0.03% |
|
Small RNA: |
8 |
1,397 bp |
0.01% |
||
|
Satellites: |
0 |
0 bp |
0.00% |
|
Satellites: |
0 |
0 bp |
0.00% |
||
|
Simple repeats: |
0 |
0 bp |
0.00% |
|
Simple repeats: |
0 |
0 bp |
0.00% |
||
|
Low complexity: |
0 |
0 bp |
0.00% |
|
Low complexity: |
0 |
0 bp |
0.00% |
||
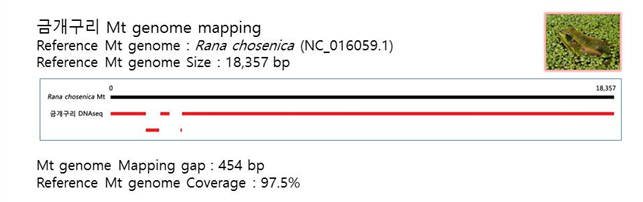
D. Mito-genome Map

(4) Microsatellite 후보군
A. SSR statistics
|
Motif(-mer) |
2 |
3 |
4 |
5 |
6 |
7 |
8 |
9 |
10 |
Total |
|
금개구리 |
556 |
68 |
22 |
3 |
- |
- |
- |
- |
- |
649 |
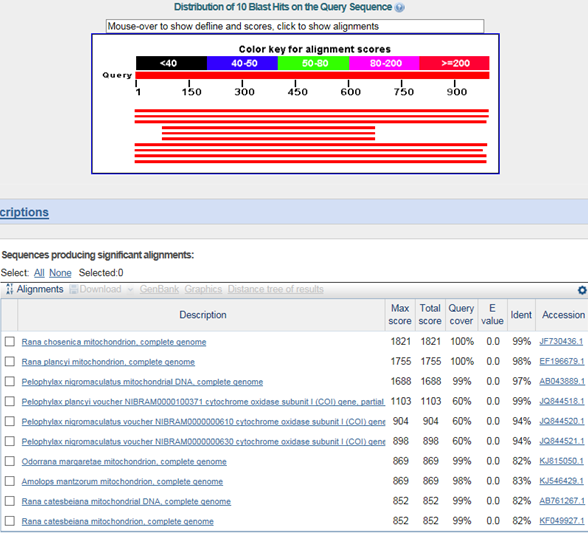
B. 바코드서열
>금개구리(Pelophylax chosenicus)
ATATTTACCCGTTGATTCTTCTCTACTAACCATAAAGATATCGGAACCCTTTACTTAATC
TTTGGCGCATGAGCAGGGATAGTCGGCACAGCCTTAAGCCTGCTTATCCGAGCGGAATTA
AGCCAACCCGGAACCCTTCTCGGCGATGACCAAATCTACAACGTAATTGTTACCGCCCAC
GCTTTTGTAATAATTTTCTTCATAGTCATGCCTATTCTAATCGGGGGCTTCGGAAACTGA
CTTGTCCCACTAATGATCGGCGCCCCTGACATGGCCTTCCCACGAATAAACAATATAAGC
TTCTGGCTTCTACCACCCTCCTTCTTTCTTCTCCTAGCTTCCTCGACAGTAGAAGCAGGA
GCAGGGACAGGTTGAACTGTTTACCCCCCTCTAGCCGGCAATCTCGCCCATGCCGGACCG
TCTGTAGACCTGGCCATTTTTTCCCTTCACTTAGCCGGGGTTTCATCAATTTTAGGGGCT
ATTAATTTTATTACAACTATTATTAATATGAAACCCACATCTATTACACAATACCAAACA
CCCCTATTCGTTTGATCTGTATTAATTACCGCCGTACTACTTCTGCTCTCCCTTCCTGTA
CTGGCAGCCGGCATTACAATACTTCTTACTGACCGTAATCTAAATACAACATTCTTTGAC
CCAGCGGGTGGAGGAGATCCCATCCTTTACCAACACTTATTCTGATTCTTTGGCCATCCT
GAAGTCTACATTCTGATCCTCCCAGGATTCGGAATTATTTCCCACGTAGTAGCCTACTAC
TCCAACAAAAAAGAACCTTTTGGTTATATGGGCATGGTCTGAGCAATATTATCCATTGGC
CTCTTGGGCTTTATCGTGTGGGCTCATCATATATTTACCACAGACCTTAATGTAGACACA
CGCGCCTATTTTACATCAGCAACAATAATCATTGCCATCCCAACCGGAGTCAAAGTATTT
AGCTGACTGGCCACCATGCACGGAGGAATCATCAA