АэПюБюИЗГыЗЁБт (Oxidus gracilis)
|
|
ЂУ БЙ Иэ: АэПюБюИЗГыЗЁБт ЂУ Ча Иэ: Oxidus gracilis ЂУ СіСЄЙјШЃ: АэРЏСО ЂУ Аш Хы: ГыЗЁБтА-ЖьГыЗЁБтИё-ЙЋДчГыЗЁБтАњ |
Ён Л§БшЛѕ : ИіРК ШцЛіРЬГЊ АЅЛіРЬАэ ИіХыГЏАГ, ВПИЎ, УЫАЂРЧ ГЁ, ДйИЎ, Йш ТЪРЧ ЛіБђРК ШВЙщЛіРЬДй. ИіБцРЬДТ 24~25mmРЬДй. ИіХы ДмИщРК ЕеБлАэ УјИщПЁ ИіХы ГЏАГАЁ ЕкТЪРИЗЮ ЛЯСЗЧЯАд ГЊ РжДй. ИіХыГЏАГДТ 3ЙјТА ИіИЖЕ№КЮХЭ ЙпДоЧЯПЉ 7ЙјТАБюСі ФПСіДйАЁ РЬШФКЮХЭ ЧіРњЧЯАд РлОЦСЎ 19ЙјТА ИЖЕ№ПЁМ ЛчЖѓСіДТ ЦЏТЁРЛ КИРЮДй. ИіХыРК Ею ТЪПЁМ КИИщ 3ЙјТА ИЖЕ№БюСіДТ СЁТї СМОЦСіДйАЁ ДйНУ ГаОюСЎ 15~16ЙјТА ИіИЖЕ№КЮХЭ СЁТї СМОЦСјДй. МіФЦРЧ 4ЙјТА ЙшЦЧПЁ ЛяАЂЧќ И№ОчРЧ ИЗСЖАЂРЬ ОјДй. МіФЦРЧ Л§НФСіДТ БтР§Ањ РќХ№Р§РЬ ТЊАэ МОХаРЬ ЙаЛ§ЧбДй. Х№Р§РЧ ИЛДмКЮДТ АяКРИ№ОчРЬИч АцР§РК ЦэЦђЧб УЪНТДо И№ОчРЧ АЁСіЗЮ ЙиКЮКаПЁ ФПДйЖѕ ПЗАЁСіАЁ РжДй. КЮР§РК ГаАэ ЕЮАЁСіЗЮ ГЊДЕИч ИЛДмКЮДТ ПЉЗЏ АЅЗЁЗЮ АЅЖѓСЎ КЏРЬАЁ ИЙДй. РЭЛъГыЗЁБтПЭ РЏЛчЧЯСіИИ МіФЦ Л§НФСі ИЛДмКЮАЁ ДйИЅ АЭРИЗЮ БИКаЧв Мі РжДй.
Ён МНФСі : РЮАЁ КЮБйПЁ МНФЧЯИч НтРК ТЄРЬГЊ КИИЎТЄПЁМ 5~9Пљ ДыЙпЛ§РЛ ЧЯБтЕЕ ЧбДй.
Ён КаЦї : ЧбБЙРЬ И№НФЛъСіЗЮ УпСЄЕЧДТ ГыЗЁБтЗЮ РЏРЯЧЯАд Рќ ММАшПЁ КаЦїЧЯДТ СОРЬДй.
1) ӪѧѪКИ
- УЄС§ЧуАЁ: (АэРЏСО)
- УЄ С§ Рк: АцКЯДыЧаБГ ШВРЧПэ
2) УЄС§ЧіШВ Йз ЛљЧУУГИЎ
|
АэПюБюИЗГыЗЁБт УЄС§ЧіШВ |
ЛљЧУУГИЎ |
|||
|
УЄС§РЯ |
АГУМ Мі |
DNAseq |
RNAseq |
ЧЅКЛ |
|
2014.07.02 |
20 EA |
10 EA |
9 EA |
1 EA |
3) QC Data
|
ЛљЧУСО |
DNA QC |
RNA QC |
Result |
||
|
Concentration (ug/ul) |
Total Conc. (ug) |
Concentration (ug/ul) |
RIN (>7) |
||
|
АэПюБюИЗГыЗЁБт |
38.3 |
8 |
394 |
7.1 |
pass |
4) БтСИ NCBI Taxonomy РЏРќСЄКИ ЧіШВ
|
РЏРќСЄКИ СОЗљ |
Oxidus Мг РЏРќРкПј |
Oxidus gracilis РЏРќРкПј |
|
Nucleotide |
56 |
32 |
|
Protein |
34 |
27 |
5) Sequencing КаМЎ АсАњ
- Illumina DNA Sequncing АсАњ
|
Input information |
|||||
|
ЁЁ |
ЁЁ |
ЁЁ |
Region1 |
Region2 |
Total |
|
raw data |
|||||
|
Number Of Reads |
|
|
144,946,656 |
144,946,656 |
289,893,312 |
|
Number Of Bases |
|
|
18,263,278,656 |
18,263,278,656 |
36,526,557,312 |
|
error correction |
|||||
|
Number Of Reads |
|
|
139,955,794 |
138,060,685 |
278,016,479 |
|
Number Of Bases |
|
|
17,182,275,706 |
16,976,750,298 |
34,159,026,004 |
|
pair reads |
|||||
|
Number Of Reads |
|
|
|
|
272,536,504 |
|
Number Of Bases |
|
|
|
|
33,568,378,377 |
|
single reads |
|||||
|
Number Of Reads |
|
|
|
|
5,479,975 |
|
Number Of Bases |
|
|
|
|
590,647,627 |
|
Assembly Results |
|||||
|
Scaffold Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Scaffolds |
894,760 |
894,760 |
88,093 |
23,931 |
5,509 |
|
Number Of Bases |
234,980,039 |
234,980,039 |
90,213,525 |
46,600,245 |
22,268,061 |
|
Avg. Scaffold Size |
262 |
262 |
1,024 |
1,947 |
4,042 |
|
N50 Scaffold Size |
343 |
343 |
1,031 |
1,907 |
4,041 |
|
N80 Scaffold Size |
141 |
141 |
648 |
1,225 |
2,513 |
|
N90 Scaffold Size |
118 |
118 |
568 |
1,101 |
2,220 |
|
largest Scaffold Size |
106,709 |
106,709 |
106,709 |
106,709 |
106,709 |
|
Contig Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Contigs |
5,973,946 |
1,277,009 |
55,077 |
9,797 |
1,293 |
|
Number Of Bases |
554,231,917 |
248,936,410 |
46,751,341 |
16,613,261 |
5,632,858 |
|
Avg. Contig Size |
92 |
194 |
848 |
1,695 |
4,356 |
|
N50 Contig Size |
91 |
191 |
814 |
1,522 |
4,643 |
|
N80 Contig Size |
58 |
125 |
592 |
1,145 |
2,468 |
|
N90 Contig Size |
49 |
112 |
542 |
1,067 |
2,200 |
|
Largest Contig Size |
84,390 |
84,390 |
84,390 |
84,390 |
84,390 |
- Illumina RNA Sequncing АсАњ
|
Input information |
|||||
|
ЁЁ |
ЁЁ |
ЁЁ |
Region1 |
Region2 |
Total |
|
raw data |
|||||
|
Number Of Reads |
|
|
122,736,242 |
122,736,242 |
245,472,484 |
|
Number Of Bases |
|
|
15,464,766,492 |
15,464,766,492 |
30,929,532,984 |
|
error correction |
|||||
|
Number Of Reads |
|
|
121,738,700 |
121,211,274 |
242,949,974 |
|
Number Of Bases |
|
|
15,038,386,396 |
14,994,658,509 |
30,033,044,905 |
|
pair reads |
|||||
|
Number Of Reads |
|
|
|
|
241,445,970 |
|
Number Of Bases |
|
|
|
|
29,873,484,731 |
|
single reads |
|||||
|
Number Of Reads |
|
|
|
|
1,504,004 |
|
Number Of Bases |
|
|
|
|
159,560,174 |
|
Assembly Results |
|||||
|
Scaffold Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Scaffolds |
156,950 |
156,950 |
26,043 |
9,897 |
2,401 |
|
Number Of Bases |
54,556,885 |
54,556,885 |
28,788,025 |
17,640,506 |
7,420,349 |
|
Avg. Scaffold Size |
347 |
347 |
1,105 |
1,782 |
3,090 |
|
N50 Scaffold Size |
548 |
548 |
1,224 |
1,788 |
2,983 |
|
N80 Scaffold Size |
187 |
187 |
711 |
1,246 |
2,287 |
|
N90 Scaffold Size |
139 |
139 |
595 |
1,112 |
2,138 |
|
largest Scaffold Size |
18,844 |
18,844 |
18,844 |
18,844 |
18,844 |
|
Contig Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Contigs |
1,872,130 |
215,119 |
21,276 |
6,251 |
1,073 |
|
Number Of Bases |
163,855,761 |
56,781,507 |
20,109,034 |
9,864,065 |
2,938,155 |
|
Avg. Contig Size |
87 |
263 |
945 |
1,577 |
2,738 |
|
N50 Contig Size |
91 |
317 |
984 |
1,550 |
2,618 |
|
N80 Contig Size |
53 |
153 |
641 |
1,171 |
2,210 |
|
N90 Contig Size |
47 |
127 |
564 |
1,081 |
2,089 |
|
Largest Contig Size |
10,342 |
10,342 |
10,342 |
10,342 |
10,342 |
(1) АэЧАСњ genomic DNA УЄУы Йз ЧЅКЛШЎКИ
|
АЫМіПы ЛљЧУ DNA ГѓЕЕ Йз Уб Оч, ШЎСѕЧЅКЛ ЧіШВЧЅ |
|||||||
|
ЁЁSampleИэ |
ГѓЕЕ(ng/ul) |
Volume(ul) |
DNA УбЗЎ(ug) |
O.D A260/280 |
ШЎСѕЧЅКЛ |
УЄС§ЧуАЁРЯ |
УЄС§РЯ |
|
АэПюБюИЗГыЗЁБт |
35 |
996 |
35 |
1.3 |
Ёл |
АэРЏСО |
2014.7.2 |
(2) РЏРќУМ МПШЎКИ : ЧіРч 20СО ШЎКИ
|
Sample |
БИКа |
raw data size |
trim data size |
КёАэ |
|
АэПюБюИЗГыЗЁБт |
DNA |
36,526,557,312 |
34,159,026,004 |
ЁЁ |
|
RNA |
30,929,532,984 |
30,033,044,905 |
ЁЁ |
(3) БтХИКаМЎ
A. Genome Size estimation
|
Лљ ЧУ Иэ |
Kmer size |
Kmer total |
Pk depth |
Genome_size |
Coverage(X) |
|
АэПюБюИЗГыЗЁБт |
23 |
22,611,678,336 |
76 |
297,522,083 |
97.4359 |
B. ЙнКЙМП
- НУФіНЬРЬ ПЯЗсЕШ АэПюБюИЗГыЗЁБт DNA Йз RNAРЧ contigs Сп 1kb РЬЛѓРЧ contigsРЧ ЙнКЙМПРЛ ХНЛі(Left; Repeat masking results of DNA contigs, Right; Repeat masking results of RNA contigs).
|
sequences: 9,797 |
|
sequences: 6.251 |
||||||||
|
total length: 16,613,261 bp (16,613,261 bp excl N/X-runs) |
|
total length: 9,864,065 bp (9,864,065 bp excl N/X-runs) |
||||||||
|
GC level: 36.43% |
|
GC level: 42.95% |
||||||||
|
bases masked: 10,066 bp (0.06%) |
|
bases masked: 3,433 bp (0.03%) |
||||||||
|
|
number of elements |
length occupied |
percentage of sequence |
|
|
number of elements |
length occupied |
percentage of sequence |
||
|
SINEs: |
1 |
30 bp |
0.00% |
|
SINEs: |
0 |
0 bp |
0.00% |
||
|
|
ALUs |
0 |
0 bp |
0.00% |
|
|
ALUs |
0 |
0 bp |
0.00% |
|
|
MIRs |
0 |
0 bp |
0.00% |
|
|
MIRs |
0 |
0 bp |
0.00% |
|
LINEs: |
7 |
390 bp |
0.00% |
|
LINEs: |
4 |
214 bp |
0.00% |
||
|
|
LINE1 |
1 |
58 bp |
0.00% |
|
|
LINE1 |
0 |
0 bp |
0.00% |
|
|
LINE2 |
0 |
0 bp |
0.00% |
|
|
LINE2 |
0 |
0 bp |
0.00% |
|
|
L3/CR1 |
2 |
153 bp |
0.00% |
|
|
L3/CR1 |
3 |
157 bp |
0.00% |
|
LTR elements: |
4 |
450 bp |
0.00% |
|
LTR elements: |
4 |
252 bp |
0.00% |
||
|
|
ERVL |
0 |
0 bp |
0.00% |
|
|
ERVL |
0 |
0 bp |
0.00% |
|
|
ERVL-MaLRs |
0 |
0 bp |
0.00% |
|
|
ERVL-MaLRs |
0 |
0 bp |
0.00% |
|
|
ERV_classI |
0 |
0 bp |
0.00% |
|
|
ERV_classI |
1 |
47 bp |
0.00% |
|
|
ERV_classII |
0 |
0 bp |
0.00% |
|
|
ERV_classII |
0 |
0 bp |
0.00% |
|
DNA elements: |
15 |
3,152 bp |
0.02% |
|
DNA elements: |
10 |
1,495 bp |
0.02% |
||
|
|
hAT-Charlie |
5 |
1,155 bp |
0.01% |
|
|
hAT-Charlie |
5 |
829 bp |
0.01% |
|
|
TcMar-Tigger |
5 |
1,329 bp |
0.01% |
|
|
TcMar-Tigger |
1 |
52 bp |
0.00% |
|
Unclassified: |
0 |
0 bp |
0.00% |
|
Unclassified: |
1 |
188 bp |
0.00% |
||
|
Total interspersed repeats: |
|
4,022 bp |
0.02% |
|
Total interspersed repeats: |
|
2,149 bp |
0.02% |
||
|
Small RNA: |
60 |
6,044 bp |
0.04% |
|
Small RNA: |
14 |
1,284 bp |
0.01% |
||
|
Satellites: |
0 |
0 bp |
0.00% |
|
Satellites: |
0 |
0 bp |
0.00% |
||
|
Simple repeats: |
0 |
0 bp |
0.00% |
|
Simple repeats: |
0 |
0 bp |
0.00% |
||
|
Low complexity: |
0 |
0 bp |
0.00% |
|
Low complexity: |
0 |
0 bp |
0.00% |
||
D. Mito-genome Map
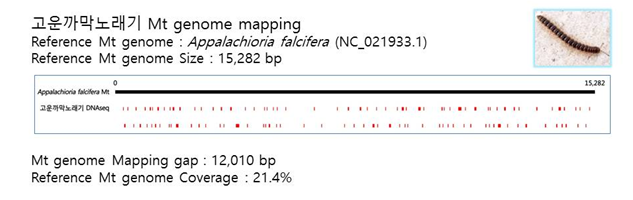
(4) Microsatellite ШФКИБК
A. SSR statistics
|
Motif(-mer) |
2 |
3 |
4 |
5 |
6 |
7 |
8 |
9 |
10 |
Total |
|
АэПюБюИЗГыЗЁБт |
461 |
272 |
20 |
4 |
5 |
2 |
- |
- |
2 |
766 |
B. ЙйФкЕхМП
ОјРН.
