염주알다슬기 (Koreanomelania nodifira)
|
|
▣ 국 명: 염주알다슬기 ▣ 학 명: Koreanomelania nodifira ▣ 지정번호: 멸종위기동물Ⅱ급 ▣ 계 통: 복족강-흡강목-다슬기과 |
◎ 생김새 : 복족류에 속하며 껍데기가 염주 모양으로 생긴 데서 이름이 유래했다. 껍데기 높이 20mm, 지금 13mm이다. 서식지가 물살이 매우 빠른 곳이기 때문에 이런 환경에 적응하기 위하여 껍데기는 염주처럼 둥글고 매끄럽고 다른 다슬기에 비해 거의 2배 정도의 넓은 발을 갖는다. 껍데기 빛깔은 서식지에 따라 황록색?흑갈색?적갈색을 띤다. 3층의 나층을 가지고 있으며 각정은 없고 체층만 남아있는 것이 특징이다. 개체에 따라 체층에 7~8개의 나륵을 가지고 있는 경우도 있으며, 성장선이 뚜렷한 모양을 이루는 개체도 있다. 껍데기 주둥이는 크고 둥글고 체증 끝의 성장맥이 보인다.
◎ 서식지 : 수심이 약간 깊은 하천 상류의 유속이 빠른 지역에 서식하며 서식지에 따라 각피의 색이 황록색, 흑갈색, 적갈색 등을 띈다.
◎ 먹이 : 주로 녹조류나 플랑크톤, 물이끼 등을 먹고 산다.
◎ 번식 : 난생을 하며 수컷은 오른쪽 두족 부분에 산란홈이 있고 암컷은 발 부분에 산란홈을 갖는다. 번식기나 번식 행동에 대해 알려진 바는 없다.
◎ 분포 : 한국의 강원도?충청북도 등 중부지방에 분포한다. 문경군 마성면 구랑, 회양, 남한강, 북한강, 전라도에서 채집된다.
1) 채집정보
- 채집허가: 2014.06.30 / 5마리
- 채 집 자: 강원대학교 이준상
2) 채집현황 및 샘플처리
|
염주알다슬기 채집현황 |
샘플처리 |
|||
|
채집일 |
개체 수 |
DNAseq |
RNAseq |
표본 |
|
2014.07.31 |
5 EA |
2 EA |
2 EA |
1 EA, 70% EtOH |
3) QC Data
|
샘플종 |
DNA QC |
RNA QC |
Result |
||
|
Concentration (ug/ul) |
Total Conc. (ug) |
Concentration (ug/ul) |
RIN (>7) |
||
|
염주알다슬기 |
202.1 |
16.2 |
772.4 |
10 |
pass |
4) 기존 NCBI Taxonomy 유전정보 현황
|
유전정보 종류 |
Koreanomelania 속 유전자원 |
Koreanomelania nodifira 유전자원 |
|
Nucleotide |
32 |
31 |
|
Protein |
16 |
15 |
5) Sequencing 분석 결과
- Illumina DNA Sequncing 결과
|
Input information |
|||||
|
|
|
|
Region1 |
Region2 |
Total |
|
raw data |
|||||
|
Number Of Reads |
|
|
152,042,627 |
152,042,627 |
304,085,254 |
|
Number Of Bases |
|
|
19,157,371,002 |
19,157,371,002 |
38,314,742,004 |
|
error correction |
|||||
|
Number Of Reads |
|
|
148,380,867 |
146,356,555 |
294,737,422 |
|
Number Of Bases |
|
|
18,115,869,900 |
17,747,931,613 |
35,863,801,513 |
|
pair reads |
|||||
|
Number Of Reads |
|
|
|
|
289,855,002 |
|
Number Of Bases |
|
|
|
|
35,336,376,873 |
|
single reads |
|||||
|
Number Of Reads |
|
|
|
|
4,882,420 |
|
Number Of Bases |
|
|
|
|
527,424,640 |
|
Assembly Results |
|||||
|
Scaffold Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Scaffolds |
1,532,046 |
1,532,046 |
435,275 |
147,299 |
26,307 |
|
Number Of Bases |
683,896,137 |
683,896,137 |
435,632,051 |
234,861,268 |
72,439,002 |
|
Avg. Scaffold Size |
446 |
446 |
1,000 |
1,594 |
2,753 |
|
N50 Scaffold Size |
697 |
697 |
1,061 |
1,574 |
2,654 |
|
N80 Scaffold Size |
302 |
302 |
652 |
1,182 |
2,201 |
|
N90 Scaffold Size |
186 |
186 |
586 |
1,085 |
2,091 |
|
largest Scaffold Size |
26,917 |
26,917 |
26,917 |
26,917 |
26,917 |
|
Contig Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Contigs |
39,252,622 |
2,718,564 |
265,405 |
36,135 |
3,108 |
|
Number Of Bases |
2,186,714,551 |
676,568,570 |
201,823,280 |
50,238,238 |
8,267,642 |
|
Avg. Contig Size |
55 |
248 |
760 |
1,390 |
2,660 |
|
N50 Contig Size |
55 |
317 |
736 |
1,321 |
2,515 |
|
N80 Contig Size |
33 |
147 |
575 |
1,096 |
2,159 |
|
N90 Contig Size |
32 |
118 |
535 |
1,044 |
2,077 |
|
Largest Contig Size |
17,042 |
17,042 |
17,042 |
17,042 |
17,042 |
- Illumina RNA Sequncing 결과
|
Input information |
|||||
|
|
|
|
Region1 |
Region2 |
Total |
|
raw data |
|||||
|
Number Of Reads |
|
|
154,130,822 |
154,130,822 |
308,261,644 |
|
Number Of Bases |
|
|
19,420,483,572 |
19,420,483,572 |
38,840,967,144 |
|
error correction |
|||||
|
Number Of Reads |
|
|
151,145,649 |
149,514,224 |
300,659,873 |
|
Number Of Bases |
|
|
18,539,686,152 |
18,323,040,456 |
36,862,726,608 |
|
pair reads |
|||||
|
Number Of Reads |
|
|
|
|
297,086,402 |
|
Number Of Bases |
|
|
|
|
36,488,385,842 |
|
single reads |
|||||
|
Number Of Reads |
|
|
|
|
3,573,471 |
|
Number Of Bases |
|
|
|
|
374,340,766 |
|
Assembly Results |
|||||
|
Scaffold Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Scaffolds |
407,270 |
407,270 |
69,731 |
27,341 |
8,533 |
|
Number Of Bases |
150,948,261 |
150,948,261 |
83,308,077 |
54,037,232 |
28,212,304 |
|
Avg. Scaffold Size |
370 |
370 |
1,194 |
1,976 |
3,306 |
|
N50 Scaffold Size |
600 |
600 |
1,371 |
2,074 |
3,279 |
|
N80 Scaffold Size |
206 |
206 |
738 |
1,316 |
2,389 |
|
N90 Scaffold Size |
143 |
143 |
607 |
1,146 |
2,177 |
|
largest Scaffold Size |
26,699 |
26,699 |
26,699 |
26,699 |
26,699 |
|
Contig Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Contigs |
7,163,882 |
558,193 |
58,091 |
17,338 |
3,891 |
|
Number Of Bases |
432,571,802 |
150,408,573 |
57,426,119 |
29,705,264 |
11,595,857 |
|
Avg. Contig Size |
60 |
269 |
988 |
1,713 |
2,980 |
|
N50 Contig Size |
63 |
344 |
1,031 |
1,714 |
2,896 |
|
N80 Contig Size |
34 |
151 |
650 |
1,209 |
2,271 |
|
N90 Contig Size |
32 |
120 |
568 |
1,096 |
2,129 |
|
Largest Contig Size |
14,658 |
14,658 |
14,658 |
14,658 |
14,658 |
나) 멸종위기종 동물 Genome Survey
|
검수용 샘플 DNA 농도 및 총 양, 확증표본 현황표 |
|||||||
|
Sample명 |
농도(ng/ul) |
Volume(ul) |
DNA 총량(ug) |
O.D A260/280 |
확증표본 |
채집허가일 |
채집일 |
|
염주알다슬기 |
441 |
996 |
439 |
1.84 |
○ |
2014.6.30 |
2014.7.31 |
(2) 유전체 서열확보 : 현재 20종 확보
|
Sample |
구분 |
raw data size |
trim data size |
비고 |
|
염주알다슬기 |
DNA |
38,314,742,004 |
35,863,801,513 |
|
|
RNA |
38,840,967,144 |
36,862,726,608 |
|
(3) 기타분석
A. Genome Size estimation
|
샘 플 명 |
Kmer size |
Kmer total |
Pk depth |
Genome_size |
Coverage(X) |
|
염주알다슬기 |
23 |
23,718,649,812 |
9 |
2,635,405,534 |
11.5385 |
B. 반복서열
- 시퀀싱이 완료된 염주알다슬기 DNA 및 RNA의 contigs 중 1kb 이상의 contigs의 반복서열을 탐색(Left; Repeat masking results of DNA contigs, Right; Repeat masking results of RNA contigs).
|
sequences: 36,135 |
|
sequences: 17,338 |
||||||||
|
total length: 50,238,238 bp (50,238,238 bp excl N/X-runs) |
|
total length: 29,705,264 bp (29,705,264 bp excl N/X-runs) |
||||||||
|
GC level: 43.77% |
|
GC level: 49.48% |
||||||||
|
bases masked: 15,139 bp (0.03%) |
|
bases masked: 5,847 bp (0.02%) |
||||||||
|
|
number of elements |
length occupied |
percentage of sequence |
|
|
number of elements |
length occupied |
percentage of sequence |
||
|
SINEs: |
12 |
829 bp |
0.00% |
|
SINEs: |
9 |
559 bp |
0.00% |
||
|
|
ALUs |
0 |
0 bp |
0.00% |
|
|
ALUs |
0 |
0 bp |
0.00% |
|
|
MIRs |
5 |
369 bp |
0.00% |
|
|
MIRs |
5 |
332 bp |
0.00% |
|
LINEs: |
80 |
7,265 bp |
0.01% |
|
LINEs: |
29 |
2,314 bp |
0.01% |
||
|
|
LINE1 |
12 |
687 bp |
0.00% |
|
|
LINE1 |
7 |
394 bp |
0.00% |
|
|
LINE2 |
19 |
1,694 bp |
0.00% |
|
|
LINE2 |
9 |
479 bp |
0.00% |
|
|
L3/CR1 |
22 |
1,577 bp |
0.00% |
|
|
L3/CR1 |
7 |
453 bp |
0.00% |
|
LTR elements: |
13 |
1,281 bp |
0.01% |
|
LTR elements: |
6 |
542 bp |
0.00% |
||
|
|
ERVL |
2 |
105 bp |
0.00% |
|
|
ERVL |
0 |
0 bp |
0.00% |
|
|
ERVL-MaLRs |
0 |
0 bp |
0.00% |
|
|
ERVL-MaLRs |
0 |
0 bp |
0.00% |
|
|
ERV_classI |
5 |
308 bp |
0.00% |
|
|
ERV_classI |
3 |
187 bp |
0.00% |
|
|
ERV_classII |
1 |
77 bp |
0.00% |
|
|
ERV_classII |
1 |
77 bp |
0.00% |
|
DNA elements: |
35 |
3,307 bp |
0.01% |
|
DNA elements: |
14 |
1,314 bp |
0.00% |
||
|
|
hAT-Charlie |
0 |
0 bp |
0.00% |
|
|
hAT-Charlie |
1 |
71 bp |
0.00% |
|
|
TcMar-Tigger |
3 |
263 bp |
0.00% |
|
|
TcMar-Tigger |
1 |
62 bp |
0.00% |
|
Unclassified: |
3 |
205 bp |
0.00% |
|
Unclassified: |
2 |
183 bp |
0.00% |
||
|
Total interspersed repeats: |
|
12,887 bp |
0.03% |
|
Total interspersed repeats: |
|
4,912 bp |
0.02% |
||
|
Small RNA: |
35 |
2,118 bp |
0.00% |
|
Small RNA: |
6 |
984 bp |
0.00% |
||
|
Satellites: |
2 |
202 bp |
0.00% |
|
Satellites: |
0 |
0 bp |
0.00% |
||
|
Simple repeats: |
0 |
0 bp |
0.00% |
|
Simple repeats: |
0 |
0 bp |
0.00% |
||
|
Low complexity: |
0 |
0 bp |
0.00% |
|
Low complexity: |
0 |
0 bp |
0.00% |
||
D. Mito-genome Map
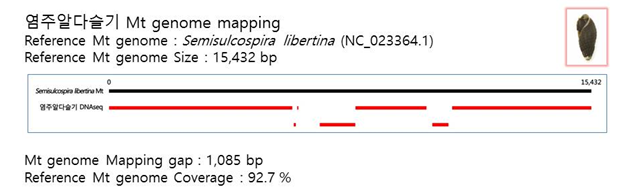
(4) Microsatellite 후보군
A. SSR statistics
|
Motif(-mer) |
2 |
3 |
4 |
5 |
6 |
7 |
8 |
9 |
10 |
Total |
|
염주알다슬기 |
7,340 |
1,503 |
507 |
71 |
11 |
- |
- |
- |
- |
9,432 |
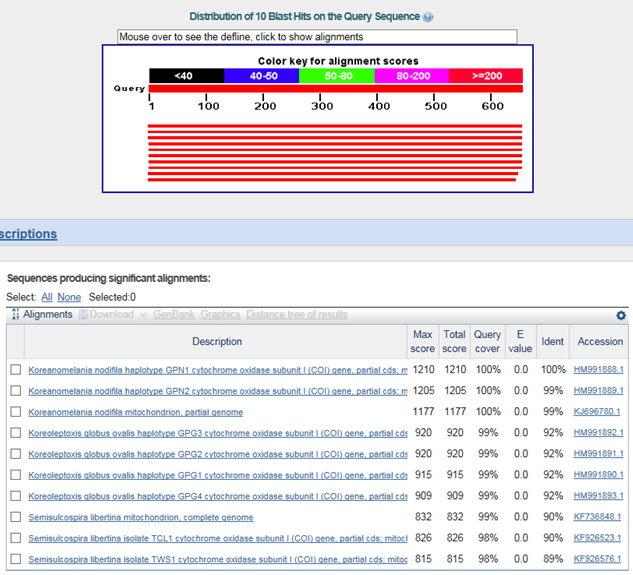
B. 바코드서열
>염주알다슬기(Koreanomelania nodifira)
TACACTTTATATTTTATTTGGAATATGATCAGGACTAGTTGGAACTGCTTTAAGCCTTTT
AATTCGTGCAGAATTAGGGCAACCAGGAGCTTTACTTGGGGATGACCAATTATATAATGT
AATTGTTACAGCCCATGCTTTCGTTATAATTTTTTTCTTAGTTATGCCCATAATAATTGG
AGGTTTTGGTAATTGACTAATTCCTTTAATGCTCGGGGCACCAGATATAGCTTTCCCGCG
CTTAAATAATATAAGTTTTTGGCTTTTACCCCCAGCACTATTGCTTTTACTCTCTTCGGC
GGCTGTCGAAAGAGGTGTAGGAACAGGGTGAACTGTTTACCCCCCTTTATCGAGAAATCT
CGCTCACGCCGGTGGCTCTGTAGATTTAGCCATCTTCTCGCTTCACTTAGCAGGTGCTTC
CTCTATTTTAGGTGCTGTAAATTTCATTACAACTATTATTAATATACGATGACGAGGAAT
ACAGTTCGAACGACTTCCTTTATTTGTTTGATCTGTAAAAATTACCGCCATTCTTCTTTT
ACTTTCTTTGCCCGTTTTGGCAGGTGCTATTACCATATTATTAACAGATCGAAATTTTAA
CACGGCTTTCTTTGACCCAGCTGGAGGAGGGGATCCAATTCTTTATCAACATCTT