북한산달팽이 (Koreanohadra (Fruticicola) kurodana)
|
|
▣ 국 명: 북한산달팽이 ▣ 학 명: Koreanohadra kurodana ▣ 지정번호: 고유종 ▣ 계 통: 복족강-진유폐목-달팽이과 |
◎ 생김새 : 패각은 낮은 구형이고 낮은 원뿔형 나탑을 갖는다. 껍질은 단단하지만 두껍지는 않으며 체색은 연갈색이고 패각 아랫면 제공과 봉합 주변은 넓게 적갈색을 띤다. 체층 주연에는 밤색의 띠가 둘러져 있고, 이후 나층에는 봉합을 따라 좁은 띠로 나타난다. 패각 표면에는 촘촘하고 불규칙한 줄 모양의 성장맥이 나선을 따라 나타나지만 첫 번째 체층은 매끄럽다. 나층은 5.5층이고 비교적 잘 부풀어 있으며 각 나층은 규칙적으로 증가한다. 껍데기 높이는 21mm, 지름은 28.6mm이고 체층의 높이가 18.5mm로 껍데기 높이의 88%를 차지한다. 봉합은 비교적 깊은 편이며 각구는 비스듬하고 넓은 반달형이다. 각구 가장자리는 얇고 퍼지며 좁은 주름을 이루고 있다. 주연의 색은 갈색을 띠고 끝 부분은 색이 흐려지는데 각축 쪽으로는 백색이고 안으로 약간 말려있는 것이 특징이다.
◎ 서식지 : 숲 속에서 습기가 높은 관목림의 낙엽 밑이나 나무 위, 응달의 돌무덤에서 서식한다.
◎ 분포 : 한국 고유종으로 서울 근교 북한산이 모식산지이며 북한산, 강원도의 화천, 춘천, 태백 등지에 분
1) 채집정보
- 채집허가: (고유종)
- 채 집 자: 강원대학교 이준상
2) 채집현황 및 샘플처리
|
북한산달팽이 채집현황 |
샘플처리 |
|||
|
채집일 |
개체 수 |
DNAseq |
RNAseq |
표본 |
|
2014.08.18 |
1 EA |
1 EA (발) |
1 EA (내장) |
- |
3) QC Data
|
샘플종 |
DNA QC |
RNA QC |
Result |
||
|
Concentration (ug/ul) |
Total Conc. (ug) |
Concentration (ug/ul) |
RIN (>7) |
||
|
북한산달팽이 |
64.7 |
7.2 |
334 |
9.3 |
pass |
4) 기존 NCBI Taxonomy 유전정보 현황
|
유전정보 종류 |
Fruticicola 속 유전자원 |
Koreanohadra (Fruticicola) kurodana 유전자원 |
|
Nucleotide |
6 |
- |
|
Protein |
11 |
- |
5) Sequencing 분석 결과
- Illumina DNA Sequncing 결과
|
Input information |
|||||
|
|
Region1 |
Region2 |
Region3 |
Region4 |
Total |
|
raw data |
|||||
|
Number Of Reads |
108,229,227 |
108,229,227 |
14,331,684 |
14,331,684 |
245,121,822 |
|
Number Of Bases |
13,636,882,602 |
13,636,882,602 |
1,805,792,184 |
1,805,792,184 |
30,885,349,572 |
|
error correction |
|||||
|
Number Of Reads |
100,857,277 |
99,362,264 |
8,460,465 |
8,246,640 |
216,926,646 |
|
Number Of Bases |
11,296,311,287 |
11,007,202,026 |
879,771,636 |
844,177,937 |
24,027,462,886 |
|
pair reads |
|||||
|
Number Of Reads |
|
187,921,310 |
|
13,248,500 |
201,169,810 |
|
Number Of Bases |
|
21,088,225,854 |
|
1,419,839,398 |
22,508,065,252 |
|
single reads |
|||||
|
Number Of Reads |
|
12,298,231 |
|
3,458,605 |
15,756,836 |
|
Number Of Bases |
|
1,215,287,459 |
|
304,110,175 |
1,519,397,634 |
|
Assembly Results |
|||||
|
Scaffold Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Scaffolds |
2,435,419 |
2,435,419 |
233,347 |
28,960 |
799 |
|
Number Of Bases |
604,222,642 |
604,222,642 |
172,374,582 |
36,718,333 |
1,863,639 |
|
Avg. Scaffold Size |
248 |
248 |
738 |
1,267 |
2,332 |
|
N50 Scaffold Size |
305 |
305 |
726 |
1,226 |
2,264 |
|
N80 Scaffold Size |
150 |
150 |
573 |
1,072 |
2,073 |
|
N90 Scaffold Size |
124 |
124 |
534 |
1,034 |
2,031 |
|
largest Scaffold Size |
5,233 |
5,233 |
5,233 |
5,233 |
5,233 |
|
Contig Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Contigs |
26,870,604 |
3,517,285 |
42,066 |
1,445 |
48 |
|
Number Of Bases |
1,974,099,305 |
649,678,319 |
27,023,646 |
1,774,873 |
113,310 |
|
Avg. Contig Size |
73 |
184 |
642 |
1,228 |
2,360 |
|
N50 Contig Size |
68 |
187 |
619 |
1,159 |
2,227 |
|
N80 Contig Size |
48 |
140 |
537 |
1,052 |
2,078 |
|
N90 Contig Size |
46 |
119 |
517 |
1,024 |
2,039 |
|
Largest Contig Size |
4,234 |
4,234 |
4,234 |
4,234 |
4,234 |
- Illumina RNA Sequncing 결과
|
Input information |
|||||
|
|
Region1 |
Region2 |
Region3 |
Region4 |
Total |
|
raw data |
|||||
|
Number Of Reads |
101,105,306 |
101,105,306 |
18,272,400 |
18,272,400 |
238,755,412 |
|
Number Of Bases |
12,739,268,556 |
12,739,268,556 |
2,302,322,400 |
2,302,322,400 |
30,083,181,912 |
|
error correction |
|||||
|
Number Of Reads |
100,049,269 |
99,293,551 |
17,547,787 |
17,386,199 |
234,276,806 |
|
Number Of Bases |
12,388,781,146 |
12,313,638,019 |
2,142,140,358 |
2,116,302,279 |
28,960,861,802 |
|
pair reads |
|||||
|
Number Of Reads |
|
197,750,080 |
|
34,302,856 |
232,052,936 |
|
Number Of Bases |
|
24,530,475,091 |
|
4,196,986,416 |
28,727,461,507 |
|
single reads |
|||||
|
Number Of Reads |
|
1,592,740 |
|
631,130 |
2,223,870 |
|
Number Of Bases |
|
171,944,074 |
|
61,456,221 |
233,400,295 |
|
Assembly Results |
|||||
|
Scaffold Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Scaffolds |
184,615 |
184,615 |
38,109 |
13,565 |
3,173 |
|
Number Of Bases |
72,355,633 |
72,355,633 |
40,396,365 |
23,448,116 |
9,328,171 |
|
Avg. Scaffold Size |
391 |
391 |
1,060 |
1,728 |
2,939 |
|
N50 Scaffold Size |
592 |
592 |
1,152 |
1,744 |
2,822 |
|
N80 Scaffold Size |
230 |
230 |
692 |
1,227 |
2,262 |
|
N90 Scaffold Size |
160 |
160 |
588 |
1,105 |
2,125 |
|
largest Scaffold Size |
16,779 |
16,779 |
16,779 |
16,779 |
16,779 |
|
Contig Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Contigs |
2,100,216 |
251,317 |
31,062 |
8,752 |
1,650 |
|
Number Of Bases |
187,572,963 |
74,959,664 |
29,323,557 |
14,184,196 |
4,617,538 |
|
Avg. Contig Size |
89 |
298 |
944 |
1,620 |
2,798 |
|
N50 Contig Size |
91 |
371 |
971 |
1,609 |
2,666 |
|
N80 Contig Size |
51 |
176 |
635 |
1,190 |
2,222 |
|
N90 Contig Size |
47 |
142 |
561 |
1,090 |
2,103 |
|
Largest Contig Size |
10,777 |
10,777 |
10,777 |
10,777 |
10,777 |
(1) 고품질 genomic DNA 채취 및 표본확보
|
검수용 샘플 DNA 농도 및 총 양, 확증표본 현황표 |
|||||||
|
Sample명 |
농도(ng/ul) |
Volume(ul) |
DNA 총량(ug) |
O.D A260/280 |
확증표본 |
채집허가일 |
채집일 |
|
북한산달팽이 |
1190 |
996 |
1185 |
1.92 |
X |
고유종 |
2014.8.18 |
(2) 유전체 서열확보 : 현재 20종 확보
|
Sample |
구분 |
raw data size |
trim data size |
비고 |
|
북산한달팽이 |
DNA |
30,885,349,572 |
24,027,462,886 |
|
|
RNA |
30,083,181,912 |
28,960,861,802 |
|
(3) 기타분석
A. Genome Size estimation
|
샘 플 명 |
Kmer size |
Kmer total |
Pk depth |
Genome_size |
Coverage(X) |
|
북한산달팽이 |
23 |
16,883,759,412 |
3 |
5,627,919,804 |
3.84615 |
B. 반복서열
- 시퀀싱이 완료된 북한산달팽이 DNA 및 RNA의 contigs 중 1kb 이상의 contigs의 반복서열을 탐색(Left; Repeat masking results of DNA contigs, Right; Repeat masking results of RNA contigs).
|
sequences: 1,485 |
|
sequences: 7,116 |
||||||||
|
total length: 1,826,882 bp (1,826,882 bp excl N/X-runs) |
|
total length: 11,689,382 bp (11,689,382 bp excl N/X-runs) |
||||||||
|
GC level: 40.45% |
|
GC level: 45.61% |
||||||||
|
bases masked: 4,154 bp (0.23%) |
|
bases masked: 4,385 bp (0.04%) |
||||||||
|
|
number of elements |
length occupied |
percentage of sequence |
|
|
number of elements |
length occupied |
percentage of sequence |
||
|
SINEs: |
1 |
82 bp |
0.00% |
|
SINEs: |
2 |
113 bp |
0.00% |
||
|
|
ALUs |
0 |
0 bp |
0.00% |
|
|
ALUs |
0 |
0 bp |
0.00% |
|
|
MIRs |
0 |
0 bp |
0.00% |
|
|
MIRs |
1 |
58 bp |
0.00% |
|
LINEs: |
7 |
1,657 bp |
0.09% |
|
LINEs: |
17 |
994 bp |
0.01% |
||
|
|
LINE1 |
0 |
0 bp |
0.00% |
|
|
LINE1 |
0 |
0 bp |
0.00% |
|
|
LINE2 |
1 |
37 bp |
0.00% |
|
|
LINE2 |
4 |
241 bp |
0.00% |
|
|
L3/CR1 |
0 |
0 bp |
0.00% |
|
|
L3/CR1 |
6 |
343 bp |
0.00% |
|
LTR elements: |
3 |
311 bp |
0.02% |
|
LTR elements: |
3 |
324 bp |
0.00% |
||
|
|
ERVL |
0 |
0 bp |
0.00% |
|
|
ERVL |
1 |
62 bp |
0.00% |
|
|
ERVL-MaLRs |
0 |
0 bp |
0.00% |
|
|
ERVL-MaLRs |
0 |
0 bp |
0.00% |
|
|
ERV_classI |
3 |
311 bp |
0.02% |
|
|
ERV_classI |
1 |
67 bp |
0.00% |
|
|
ERV_classII |
0 |
0 bp |
0.00% |
|
|
ERV_classII |
0 |
0 bp |
0.00% |
|
DNA elements: |
11 |
1,989 bp |
0.11% |
|
DNA elements: |
22 |
2,523 bp |
0.02% |
||
|
|
hAT-Charlie |
3 |
298 bp |
0.02% |
|
|
hAT-Charlie |
9 |
638 bp |
0.01% |
|
|
TcMar-Tigger |
0 |
0 bp |
0.01% |
|
|
TcMar-Tigger |
5 |
532 bp |
0.00% |
|
Unclassified: |
0 |
0 bp |
0.00% |
|
Unclassified: |
0 |
0 bp |
0.00% |
||
|
Total interspersed repeats: |
|
4,039 bp |
0.22% |
|
Total interspersed repeats: |
|
3,954 bp |
0.03% |
||
|
Small RNA: |
1 |
115 bp |
0.01% |
|
Small RNA: |
4 |
388 bp |
0.00% |
||
|
Satellites: |
0 |
0 bp |
0.00% |
|
Satellites: |
1 |
98 bp |
0.00% |
||
|
Simple repeats: |
0 |
0 bp |
0.00% |
|
Simple repeats: |
0 |
0 bp |
0.00% |
||
|
Low complexity: |
0 |
0 bp |
0.00% |
|
Low complexity: |
0 |
0 bp |
0.00% |
||
D. Mito-genome Map
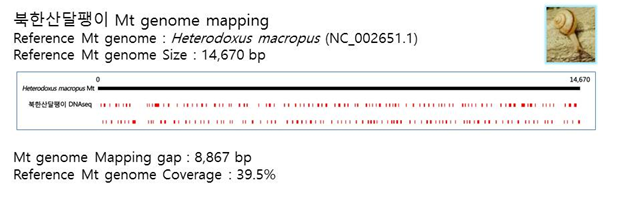

(4) Microsatellite 후보군
A. SSR statistics
|
Motif(-mer) |
2 |
3 |
4 |
5 |
6 |
7 |
8 |
9 |
10 |
Total |
|
북한산달팽이 |
429 |
200 |
39 |
4 |
1 |
- |
- |
- |
- |
673 |
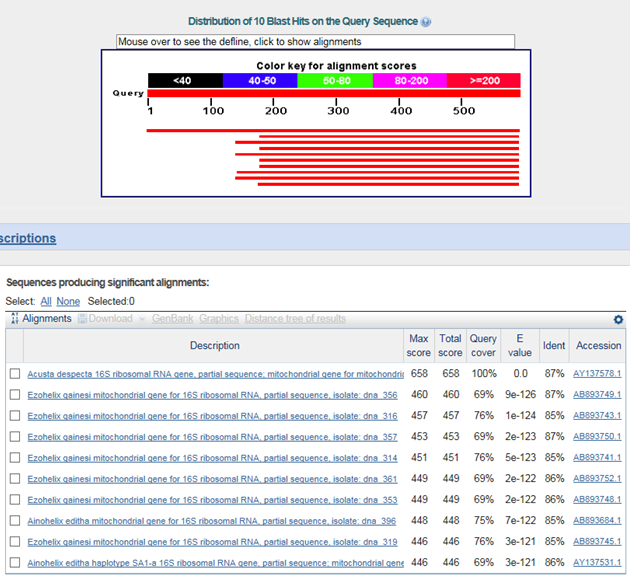
B. 바코드서열
>북한산달팽이(Koreanohadra kurodana)
ATATAGAGTGGTTAGGCTTTATATATTTATAATATTTATTTTAGAGTAATAATGATATAA
TCATCTATTATTTCATAAAATTATTATTTTAATTTAAGTTATGTATATTATATGTGGTAC
TGTATACATTTAAACCTCCGTTGATAGTGTTTTCATAAGTACTGTTGTTAGACCTAGATA
ATGAATTCGGCAAGATTTTCTTCAACTGTTTATCAAAAACATAGCTGTATGGTTGTATAT
ACAGTTTTTTCTGCTCAATGAATATAAATAGCCGCAGTACCCTGACTGTGCAAAGGTAGC
ATAATCAGTTGGCTTATAATTGAAGTCTAGTATGAACGAAAGGATGGGAGTAAGCTATCT
AAGAATTGTATTGATAAATTACTTAATAAGTGAAAATTCTTATTTAAATATAATAGACGA
GAAGACCCTAGAAATTTTAAGTAATATTTAACTATTGGTTTTTGTTGGGGCGACACAGTA
GCACTATAAACCTACTAAGAAGGATATGATTACCTTAATATGTATAAATAAATTACTCTA
GGGATAACAGCATAATATTGTAAAGTTTGTGACCTCGATGTTGGACTAGGAA
Ezohelix gainesi : 북한산달팽이 taxomony 이 확인할수 없음
Ainohelix editha : 위의 종과 genus부터 다름