ИЭВЧРЬ (Kaloula borealis)
|
|
ЂУ БЙ Иэ: ИЭВЧРЬ ЂУ ПЕ Иэ: Narrow-mouth frog ЂУ Ча Иэ: Kaloula borealis ЂУ СіСЄЙјШЃ: ИъСОРЇБтОпЛ§ЕПЙАIIБо ЂУ Аш Хы: ОчМА-ЙЋЙЬИё-ИЭВЧРЬАњ |
2) УЄС§ЧіШВ Йз ЛљЧУУГИЎ
|
ИЭВЧРЬ УЄС§ЧіШВ |
ЛљЧУУГИЎ |
|||
|
УЄС§РЯ |
АГУМ Мі |
DNAseq |
RNAseq |
ЧЅКЛ |
|
2014.07.31 |
2 EA |
2 EA (ИіХы) |
2 EA (ГЛРх) |
- |
3) QC Data
|
ЛљЧУСО |
DNA QC |
RNA QC |
Result |
||
|
Concentration (ug/ul) |
Total Conc. (ug) |
Concentration (ug/ul) |
RIN (>7) |
||
|
ИЭВЧРЬ |
87.9 |
14.6 |
631.3 |
8.1 |
pass |
4) БтСИ NCBI Taxonomy РЏРќСЄКИ ЧіШВ
|
РЏРќСЄКИ СОЗљ |
Kaloula Мг РЏРќРкПј |
Kaloula borealis РЏРќРкПј |
|
Nucleotide |
39,199 |
70 |
|
Protein |
610 |
43 |
|
Genome |
4 |
1 |
|
Gene |
100 |
13 |
5) Sequencing КаМЎ АсАњ
- Illumina DNA Sequncing АсАњ
|
Input information |
|||||
|
ЁЁ |
ЁЁ |
ЁЁ |
Region1 |
Region2 |
Total |
|
raw data |
|||||
|
Number Of Reads |
|
|
173,419,798 |
173,419,798 |
346,839,596 |
|
Number Of Bases |
|
|
21,850,894,548 |
21,850,894,548 |
43,701,789,096 |
|
error correction |
|||||
|
Number Of Reads |
|
|
169,623,481 |
167,169,797 |
336,793,278 |
|
Number Of Bases |
|
|
20,377,465,874 |
19,936,253,896 |
40,313,719,770 |
|
pair reads |
|||||
|
Number Of Reads |
|
|
|
|
328,950,756 |
|
Number Of Bases |
|
|
|
|
39,477,136,052 |
|
single reads |
|||||
|
Number Of Reads |
|
|
|
|
7,842,522 |
|
Number Of Bases |
|
|
|
|
836,583,718 |
|
Assembly Results |
|||||
|
Scaffold Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Scaffolds |
2,308,228 |
2,308,228 |
841,357 |
371,618 |
93,058 |
|
Number Of Bases |
1,307,509,700 |
1,307,509,700 |
980,113,066 |
646,613,411 |
264,134,285 |
|
Avg. Scaffold Size |
566 |
566 |
1,164 |
1,739 |
2,838 |
|
N50 Scaffold Size |
987 |
987 |
1,311 |
1,775 |
2,768 |
|
N80 Scaffold Size |
412 |
412 |
776 |
1,251 |
2,247 |
|
N90 Scaffold Size |
234 |
234 |
634 |
1,118 |
2,117 |
|
largest Scaffold Size |
23,339 |
23,339 |
23,339 |
23,339 |
23,339 |
|
Contig Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Contigs |
45,069,584 |
4,848,870 |
616,552 |
117,044 |
7,397 |
|
Number Of Bases |
2,991,458,956 |
1,336,812,483 |
496,257,814 |
157,996,323 |
17,902,981 |
|
Avg. Contig Size |
66 |
275 |
804 |
1,349 |
2,420 |
|
N50 Contig Size |
73 |
364 |
803 |
1,305 |
2,330 |
|
N80 Contig Size |
34 |
163 |
598 |
1,097 |
2,107 |
|
N90 Contig Size |
32 |
125 |
546 |
1,046 |
2,049 |
|
Largest Contig Size |
11,758 |
11,758 |
11,758 |
11,758 |
11,758 |
- Illumina RNA Sequncing АсАњ
|
Input information |
|||||
|
ЁЁ |
ЁЁ |
ЁЁ |
Region1 |
Region2 |
Total |
|
raw data |
|||||
|
Number Of Reads |
|
|
143,579,255 |
143,579,255 |
287,158,510 |
|
Number Of Bases |
|
|
18,090,986,130 |
18,090,986,130 |
36,181,972,260 |
|
error correction |
|||||
|
Number Of Reads |
|
|
141,101,937 |
139,782,733 |
280,884,670 |
|
Number Of Bases |
|
|
17,477,510,793 |
17,277,276,722 |
34,754,787,515 |
|
pair reads |
|||||
|
Number Of Reads |
|
|
|
|
278,191,056 |
|
Number Of Bases |
|
|
|
|
34,468,254,943 |
|
single reads |
|||||
|
Number Of Reads |
|
|
|
|
2,693,614 |
|
Number Of Bases |
|
|
|
|
286,532,572 |
|
Assembly Results |
|||||
|
Scaffold Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Scaffolds |
235,314 |
235,314 |
43,886 |
18,182 |
5,896 |
|
Number Of Bases |
93,094,170 |
93,094,170 |
53,961,086 |
36,194,926 |
19,288,256 |
|
Avg. Scaffold Size |
395 |
395 |
1,229 |
1,990 |
3,271 |
|
N50 Scaffold Size |
664 |
664 |
1,447 |
2,114 |
3,247 |
|
N80 Scaffold Size |
222 |
222 |
759 |
1,336 |
2,400 |
|
N90 Scaffold Size |
151 |
151 |
615 |
1,152 |
2,184 |
|
largest Scaffold Size |
20,489 |
20,489 |
20,489 |
20,489 |
20,489 |
|
Contig Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Contigs |
3,386,216 |
320,168 |
39,653 |
13,055 |
2,755 |
|
Number Of Bases |
226,095,795 |
93,751,151 |
39,980,565 |
21,724,704 |
7,825,771 |
|
Avg. Contig Size |
66 |
292 |
1,008 |
1,664 |
2,840 |
|
N50 Contig Size |
63 |
394 |
1,074 |
1,663 |
2,771 |
|
N80 Contig Size |
35 |
164 |
668 |
1,201 |
2,232 |
|
N90 Contig Size |
32 |
127 |
577 |
1,094 |
2,113 |
|
Largest Contig Size |
20,489 |
20,489 |
20,489 |
20,489 |
20,489 |
(2) РЏРќУМ МПШЎКИ : ЧіРч 20СО ШЎКИ
|
Sample |
БИКа |
raw data size |
trim data size |
КёАэ |
|
ИЭВЧРЬ |
DNA |
43,701,789,096 |
40,313,719,770 |
ЁЁ |
|
RNA |
36,181,972,260 |
34,754,787,515 |
ЁЁ |
(3) БтХИКаМЎ
A. Genome Size estimation
|
Лљ ЧУ Иэ |
Kmer size |
Kmer total |
Pk depth |
Genome_size |
Coverage(X) |
|
ИЭВЧРЬ |
23 |
27,053,488,488 |
7 |
3,864,784,069 |
8.97436 |
B. ЙнКЙМП
- НУФіНЬРЬ ПЯЗсЕШ ИЭВЧРЬ DNA Йз RNAРЧ contigs Сп 1kb РЬЛѓРЧ contigsРЧ ЙнКЙМПРЛ ХНЛі(Left; Repeat masking results of DNA contigs, Right; Repeat masking results of RNA contigs).
|
sequences: 117,044 |
|
sequences: 13.055 |
||||||||
|
total length: 157,996,323 bp (157,996,323 bp excl N/X-runs) |
|
total length: 21,724,704 bp (21,724,704 bp excl N/X-runs) |
||||||||
|
GClevel: 42.51 % |
|
GC level: 45.77% |
||||||||
|
bases masked: 241,819 bp ( 0.15%) |
|
bases masked: 17,888 bp ( 0.08 %) |
||||||||
|
|
numberof elements |
length occupied |
percentage of sequence |
|
|
numberof elements |
length occupied |
percentage of sequence |
||
|
SINEs: |
35 |
2,405 bp |
0.00% |
|
SINEs: |
0 |
0 bp |
0.00% |
||
|
|
ALUs |
0 |
0 bp |
0.00% |
|
|
ALUs |
0 |
0 bp |
0.00% |
|
|
MIRs |
13 |
590 bp |
0.00% |
|
|
MIRs |
0 |
0 bp |
0.00% |
|
LINEs: |
978 |
125,927 bp |
0.08% |
|
LINEs: |
48 |
6,260 bp |
0.03% |
||
|
|
LINE1 |
157 |
14,627 bp |
0.01% |
|
|
LINE1 |
10 |
619 bp |
0.00% |
|
|
LINE2 |
288 |
53,754 bp |
0.03% |
|
|
LINE2 |
12 |
2,708 bp |
0.01% |
|
|
L3/CR1 |
487 |
54,203 bp |
0.03% |
|
|
L3/CR1 |
22 |
2,658 bp |
0.01% |
|
LTRelements: |
243 |
58,177 bp |
0.04% |
|
LTRelements: |
20 |
6,232 bp |
0.03% |
||
|
|
ERVL |
24 |
4,921 bp |
0.00% |
|
|
ERVL |
2 |
826 bp |
0.00% |
|
|
ERVL-MaLRs |
7 |
425 bp |
0.00% |
|
|
ERVL-MaLRs |
3 |
138 bp |
0.00% |
|
|
ERV_classI |
42 |
5,859 bp |
0.00% |
|
|
ERV_classI |
8 |
1,369 bp |
0.01% |
|
|
ERV_classII |
4 |
224 bp |
0.00% |
|
|
ERV_classII |
1 |
118 bp |
0.00% |
|
DNAelements: |
496 |
40,757 bp |
0.03% |
|
DNAelements: |
25 |
3,433 bp |
0.02% |
||
|
|
hAT-Charlie |
99 |
6,093 bp |
0.00% |
|
|
hAT-Charlie |
8 |
525 bp |
0.00% |
|
|
TcMar-Tigger |
159 |
13,641 bp |
0.01% |
|
|
TcMar-Tigger |
5 |
340 bp |
0.00% |
|
Unclassified: |
31 |
2,900 bp |
0.00% |
|
Unclassified: |
1 |
51 bp |
0.00% |
||
|
Total interspersed repeats: |
|
230,166 bp |
0.15% |
|
Total interspersed repeats: |
|
15,976 bp |
0.07% |
||
|
Small RNA: |
175 |
12,965 bp |
0.01% |
|
Small RNA: |
4 |
1,912 bp |
0.01% |
||
|
Satellites: |
2 |
86 bp |
0.00% |
|
Satellites: |
0 |
0 bp |
0.00% |
||
|
Simple repeats: |
0 |
0 bp |
0.00% |
|
Simple repeats: |
0 |
0 bp |
0.00% |
||
|
Low complexity: |
0 |
0 bp |
0.00% |
|
Low complexity: |
0 |
0 bp |
0.00% |
||
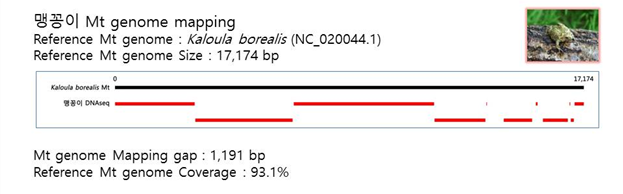
D. Mito-genome Map
4) Microsatellite ШФКИБК
A. SSR statistics
|
Motif(-mer) |
2 |
3 |
4 |
5 |
6 |
7 |
8 |
9 |
10 |
Total |
|
ИЭВЧРЬ |
9,671 |
1,324 |
477 |
64 |
5 |
- |
- |
- |
- |
11,541 |
B. ЙйФкЕхМП
>ИЭВЧРЬ(Kaloula borealis)
GGAATAGTAGGTACGGCTCTGAGCCTTCTCATTCGTGCCGAACTAAGCCAACCAGGAGCA
CTCCTAGGTGATGACCAAATTTATAATGTTATCGTTACTGCTCACGCCTTCGTAATAATT
TTCTTCATAGTTATACCCGTTATAATTGGTGGTTTTGGAAATTGACTTGTTCCCCTAATA
ATTGGAGCCCCAGACATAGCCTTCCCACGAATAAATAACATAAGCTTTTGACTTCTACCA
CCATCCTTTCTTCTTCTTTTGGCCTCATCAGCTGTAGAAGCCGGAGCCGGAACCGGCTGA
ACTGTCTACCCACCATTAGCCGGCAACCTGGCCCACGCCGGACCATCAGTTGATCTAACT
ATTTTTTCACTCCACCTGGCAGGGGTCTCTTCAATTCTTGGGGCAATTAATTTTATTACA
ACCATTTTAAATATAAAACCCCCATCAGTAACCCAATACCAAACACCCCTATTCGTCTGA
TCCGTTTTAATTACAGCTGTCCTTCTTCTTCTTTCACTCCCAGTTCTGGCAGCAGGAATT
ACAATACTGCTGACAGATCGAAACCTAAACACAACATTTTTTGACCCTGCTGGCGGTGGC