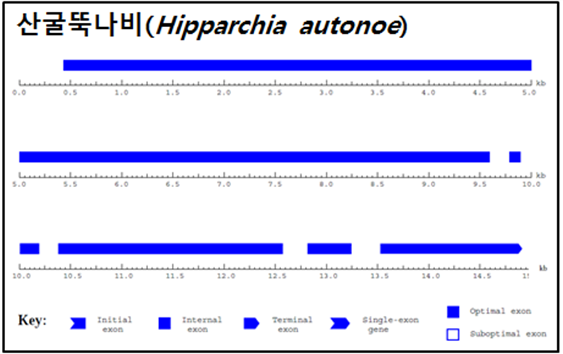
ЛъБМЖвГЊКё (Hipparchia autonoe)
|
|
ЂУ БЙ Иэ : ЛъБМЖвГЊКё ЂУ Ча Иэ : Hipparchia autonoe (Esper, 1783) ЂУ СіСЄЙјШЃ : ИъСОРЇБт ОпЛ§Л§ЙА ЅАБо ЂУ Аш Хы : ЧѳЕПЙАЙЎ > АяУцА > ГЊКёИё > ГзЙпГЊКёАњ |
Ён Чќ ХТ : ГЏАГ Цэ БцРЬДТ 45~55 mm СЄЕЕРЬДй. ИгИЎПЭ ИіРК ШцАЅЛіРЬИч, ГЏАГРЧ ЙйХСЛіЕЕ ШцАЅЛіРЛ ЖэДй. Оч ГЏАГРЧ ПмШОМБРЛ ЕћЖѓ ЙйБљТЪРИЗЮ ЙјСіДТ ШВЙщЛіРЧ ЖьЙЋДЬАЁ РжРИИч, ОеГЏАГПЁ ЙщЛіРЧ ОШСЁРЬ РжДТ 2АГРЧ ШцЛі ДЋОЫИ№Оч ЙЋДЬАЁ РжАэ, ЕоГЏАГРЧ ГЛПЌАЂ КЮБйПЁЕЕ СЖБзИЖЧб 1АГРЧ ДЋОЫИ№ОчЙЋДЬАЁ РжДй. ОЯФЦРК МіФЦПЁ КёЧЯПЉ РЯЙнРћРИЗЮ ХЉАэ ГЏАГРЧ ЙйХСЛіРЬ ПЌЧЯДй.

Ён Л§ ХТ : ГВЧбПЁМДТ ЧбЖѓЛъ 1,400 m КЮХЭ ОЯМЎРЬ ГыУтЕШ АїПЁ ЛьИч, СЄЛѓРЧ АЧСЖЧб УЪСіПЁ АГУММіАЁ РћСі ОЪДй. ЦЏШї ЧбЖѓЛъ РММПРИЇПЁМ ЙщЗЯДуПЁ РЬИЃДТ АЧСЖЧб ЧЎЙчПЁ ИЙДй. МіФЦРК ШЛъОЯ РЇПЁ ОЩОЦ НЌАэ РжРЛ ЖЇАЁ ИЙРКЕЅ, ЙйЖїРЬ ОрЧиСіИщ АэЛъСіДыРЧ МжУМВЩ, МлРЬЧЎ, ВмЧЎ ЕюРЧ ВЩПЁМ ВмРЛ КўДй. ШЛъОЯПЁМ РкСж ШоНФРЛ УыЧЯАэ ОчСіЙйИЅ АїРЧ ЧЎЙчРЬГЊ АќИё РЇИІ УЕУЕШї КёЧрЧЯДТ АЭРЛ АќТћЧв Мі РжРИИч, ЙйЖїРЬ КвИщ ИжИЎ ГЊГЊ КИХы Чб Йј ГЏОЦЕЕ 5~6 m СЄЕЕ ГЏОЦАЁ Жв ЖГОюСіЕэРЬ ГЛЗСОЩДТДй. ОЯФЦРК ИдРЬНФЙАРЧ РйПЁ ОЫРЛ Чб АГОП ГКОЦ КйРЮДй. ОжЙњЗЙЗЮ ПљЕПЧбДй.
Ён Ка Цї : БЙГЛПЁДТ СІСжЕЕ ЧбЖѓЛъ 1400 m РЬЛѓ АэСі, БЙПмПЁДТ СпБЙ, БиЕП ЗЏНУОЦ, РЏЗД Ею
Јч УЄС§ СЄКИ
- УЄ С§ Рк: КћАэРЛЛ§ХТПЌБИМв СЄЧхУЕ
- Ӫѧѳό: ЧбЖѓЛъ УЕПЌКИШЃБИПЊ РхБИИё РЯДы
Ёп ЛъБМЖвГЊКё МКУц

Ёп ЦїШЙЧб ЛъБМЖвГЊКё МКУц 4АГУМ

Јш УЄС§ЧіШВ Йз ЛљЧУУГИЎ
|
ЛъБМЖвГЊКё УЄС§ЧіШВ |
ЛљЧУУГИЎ |
|||
|
УЄС§РЯ |
АГУМ Мі |
DNAseq |
RNAseq |
ЧЅКЛ |
|
2016.07.29 |
4 EA |
2 EA |
2 EA |
- |
Јщ БтСИ NCBI Taxonomy РЏРќСЄКИ ЧіШВ
|
РЏРќСЄКИ СОЗљ |
Hipparchia Мг РЏРќРкПј |
Hipparchia autonoe РЏРќРкПј |
|
Nucleotide |
373 |
111 |
|
Protein |
360 |
96 |
|
Genome |
1 |
1 |
|
Gene |
13 |
13 |
Јъ Sequencing АсАњ
|
ЛъБМЖвГЊКё |
|||||
|
Type |
Total Reads |
Total Bases |
Total Bases (Gb) |
GC Rate |
КёАэ |
|
DNA seq |
240,779,012 |
36,357,630,812 |
36.4 |
37.2 |
|
|
RNA seq |
72,190,452 |
7,291,235,652 |
7.3 |
44.6 |
|
- Illumina DNA Sequncing АсАњ
|
SOAP de novo |
|||||
|
HA-DNA hiseq |
|||||
|
Input information |
|||||
|
ЁЁ |
ЁЁ |
ЁЁ |
Region1 |
Region2 |
Total |
|
raw data |
|||||
|
Number Of Reads |
|
|
120,389,506 |
120,389,506 |
240,779,012 |
|
Number Of Bases |
|
|
18,178,815,406 |
18,178,815,406 |
36,357,630,812 |
|
error correction |
|||||
|
Number Of Reads |
|
|
119,792,221 |
117,504,939 |
237,297,160 |
|
Number Of Bases |
|
|
17,865,202,317 |
17,052,267,774 |
34,917,470,091 |
|
pair reads |
|||||
|
Number Of Reads |
|
|
|
234,704,726 |
234,704,726 |
|
Number Of Bases |
|
|
|
34,565,893,304 |
34,565,893,304 |
|
single reads |
|||||
|
Number Of Reads |
|
|
|
2,592,434 |
2,592,434 |
|
Number Of Bases |
|
|
|
351,576,787 |
351,576,787 |
|
Assembly Results |
|||||
|
Scaffold Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1,000 bp |
> 2,000 bp |
|
Number Of Scaffolds |
|
883,724 |
115,987 |
58,729 |
30,699 |
|
Number Of Bases |
|
350,637,767 |
210,540,041 |
171,454,057 |
132,041,266 |
|
Avg. Scaffold Size |
|
396 |
1,815 |
2,919 |
4,301 |
|
N50 Scaffold Size |
|
928 |
2,897 |
3,715 |
4,715 |
|
N80 Scaffold Size |
|
198 |
1,061 |
1,843 |
2,882 |
|
N90 Scaffold Size |
|
130 |
709 |
1,382 |
2,430 |
|
largest Scaffold Size |
|
66,311 |
66,311 |
66,311 |
66,311 |
|
Contig Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1,000 bp |
> 2,000 bp |
|
Number Of Contigs |
14,640,427 |
1,384,006 |
101,536 |
24,308 |
4,271 |
|
Number Of Bases |
930,025,799 |
319,689,476 |
90,485,766 |
38,803,399 |
12,203,386 |
|
Avg. Contig Size |
63 |
230 |
891 |
1,596 |
2,857 |
|
N50 Contig Size |
66 |
266 |
892 |
1,559 |
2,733 |
|
N80 Contig Size |
36 |
135 |
609 |
1,168 |
2,219 |
|
N90 Contig Size |
34 |
114 |
550 |
1,077 |
2,103 |
|
Largest Contig Size |
14,981 |
14,981 |
14,981 |
14,981 |
14,981 |
] - Illumina RNA Sequncing АсАњ
|
Total number of raw reads |
|
|
- Number of sequences |
72,190,452 |
|
- Number of bases |
7,291,235,652 |
|
Contig information |
|
|
- Total number of contig |
161,217 |
|
- Number of bases |
162,981,105 |
|
- Mean length of contig (bp) |
1010.9 |
|
- N50 length of contig (bp) |
1,772 |
|
- GC % of contig |
40.66 |
|
- Largest contig (bp) |
28,592 |
|
- No. of large contigs (ЁУ500bp) |
84,465 |
|
Unigene information |
|
|
- Total number of unigenes |
41,602 |
|
- Number of bases |
66,264,041 |
|
- Mean length of unigene (bp) |
1592.8 |
|
- N50 length of unigene (bp) |
2,571 |
|
- GC % of unigene |
40.79 |
|
- Length ranges (bp) |
224 – 28,944 |
2) ЧіРч ЛѓШВПЁ ДыЧб АэТћ
(1) АэЧАСњ genomic DNA УЄУы Йз ЧЅКЛШЎКИ
- АЫМіПы DNAДТ ДыКЮКа ИИСЗЧЯПДАэ ИъСОРЇБтСОРЮ ДыЛѓСОЕщРК УЄС§ЗЎРЧ СІЧбРИЗЮ РЮЧи ШЎСѕЧЅКЛРИЗЮ СІУтАЁДЩЧб ЧЅКЛРЛ ШЎКИЧЯСі ИјЧЯПЉ ЛчСјРИЗЮ ДыУГЧЯАэРк Чд.
|
АЫМіПы ЛљЧУ DNA ГѓЕЕ Йз Уб Оч, ШЎСѕЧЅКЛ ЧіШВЧЅ |
||||||
|
ЁЁ |
Sample Иэ |
ГѓЕЕ(ng/ul) |
Volume(ul) |
DNA УбЗЎ(ug) |
РњРхРхМв |
ШЎСѕЧЅКЛ |
|
1 |
ЛъБМЖвГЊКё |
29.679 |
68 |
2.018 |
1XTE pH8.0 |
ЛчСјРИЗЮДыУМ |
(2) БтХИКаМЎ
A. Genome Size estimation – Jellyfish ЧСЗЮБзЗЅ
АЂ ДыЛѓСОРЛ ДыЛѓРИЗЮ Jellyfish ЧСЗЮБзЗЅРИЗЮ Genome sizeИІ ПЙУјЧб АсАњДТ ОЦЗЁПЭ ААДй. АЂ КаМЎРК k-mer АЊ 17mer, 19mer, 21merИІ ДыЛѓРИЗЮ ЧЯПДАэ БзАЭРЛ ХыЧи ПЙУјЕШ genomeРЧ ХЉБтДТ Estimation genome size ПЁ ЧЅНУЕЧОњАэ, РЬИІ ХыЧи РЬЙј ЛчОїПЁМ КаМЎЧб МПРЧ ОчРЧ coverageДТ Genome coverage depth(Coverage Depth)ПЁ ЧЅБтЕЧОњДй. БзИЎАэ АЂ СОРЧ ЧЅ ОЦЗЁ БзЗЁЧСДТ kmerАЊ КАЗЮ ГЊХИГ depthРЧ ПЙУј БзЗЁЧСЗЮ peakАЊРЛ ШЎРЮЧв Мі РжДй.
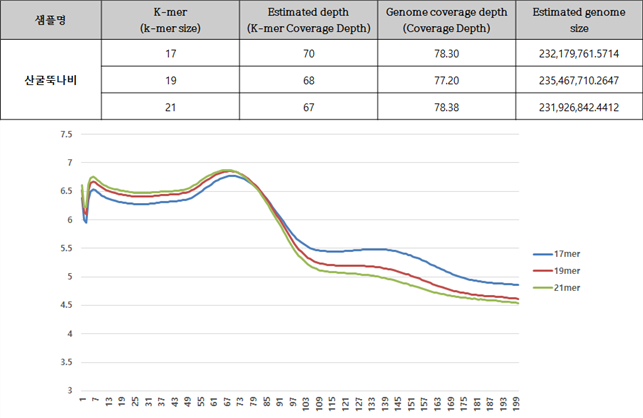
B. ЙнКЙМП
- НУФіНЬРЬ ПЯЗсЕШ ЛљЧУ DNA РЧ contigs Сп 1kb РЬЛѓРЧ contigsРЧ ЙнКЙМПРЛ ХНЛі (Repeat masking results of DNA contigs)
|
ЛъБМЖвГЊКё (Hipparchia autonoe) |
|
ЛъБМЖвГЊКё (Hipparchia autonoe) ДТ 1kb РЬЛѓРЧ contig РЧ ЙнКЙМПРК 7,838АГ read (10,187,681 bp) РЬАэ, GC level РК 32.17%РЬДй. Repeat Masking РИЗЮ ГЊПТ ЙнКЙМПРК 111,849 bp (РќУМРЧ 1.1%) ИИХ РжДй. Small RNA РК 17АГ read (733 bp) РЬ РжДй. |
||||
|
sequences: 7,838 |
|
|||||
|
total length: 10,187,681 bp (10,187,681 bp excl N/X-runs) |
|
|||||
|
GC level: 32.17 % |
|
|||||
|
bases masked: 111,849 bp ( 1.10%) |
|
|||||
|
ЁЁ |
number of elements |
length occupied |
percentage of sequence |
|
||
|
|
||||||
|
SINEs: |
8 |
526 bp |
0.00% |
|
||
|
ЁЁ |
ALUs |
0 |
0 bp |
0.00% |
|
|
|
ЁЁ |
MIRs |
5 |
318 bp |
0.00% |
|
|
|
LINEs: |
22 |
2,067 bp |
0.01% |
|
||
|
ЁЁ |
LINE1 |
6 |
302 bp |
0.00% |
|
|
|
ЁЁ |
LINE2 |
4 |
266 bp |
0.00% |
|
|
|
ЁЁ |
L3/CR1 |
8 |
442 bp |
0.00% |
|
|
|
LTRelements: |
7 |
1,089 bp |
0.00% |
|
||
|
ЁЁ |
ERVL |
1 |
59 bp |
0.00% |
|
|
|
ЁЁ |
ERVL-MaLRs |
0 |
0 bp |
0.00% |
|
|
|
ЁЁ |
ERV_classI |
1 |
97 bp |
0.00% |
|
|
|
ЁЁ |
ERV_classII |
0 |
0 bp |
0.00% |
|
|
|
DNAelements: |
35 |
4,899 bp |
0.01% |
|
||
|
ЁЁ |
hAT-Charlie |
11 |
1,430 bp |
0.00% |
|
|
|
ЁЁ |
TcMar-Tigger |
6 |
1,827 bp |
0.00% |
|
|
|
Unclassified: |
12 |
1,406 bp |
0.00% |
|
||
|
Total interspersed repeats: |
|
9,987 bp |
0.03% |
|
||
|
Small RNA: |
17 |
733 bp |
0.00% |
|
||
|
Satellites: |
5 |
290 bp |
0.00% |
|
||
C. Mito-genome Map
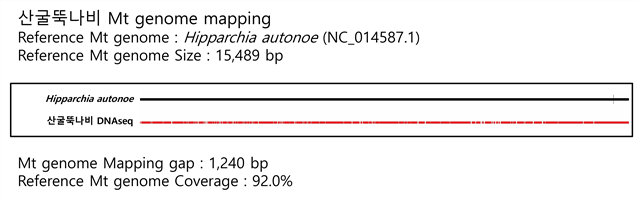
ЛъБМЖвГЊКёДТ ReferenceЗЮ 15,489 bp ХЉБтРЧ Hipparchia autonoe ЙЬХфФмЕхИЎОЦ genomeРЛ РЬПыЧЯПЉ ИЪЧЮЧЯПДДй. АИРК 1,240 bp ЗЮ, Coverage ДТ 92.0%ЗЮ ШЎРЮЕЧОњДй.
D. Transcriptome data КаМЎ
- ЛъБМЖвГЊКё
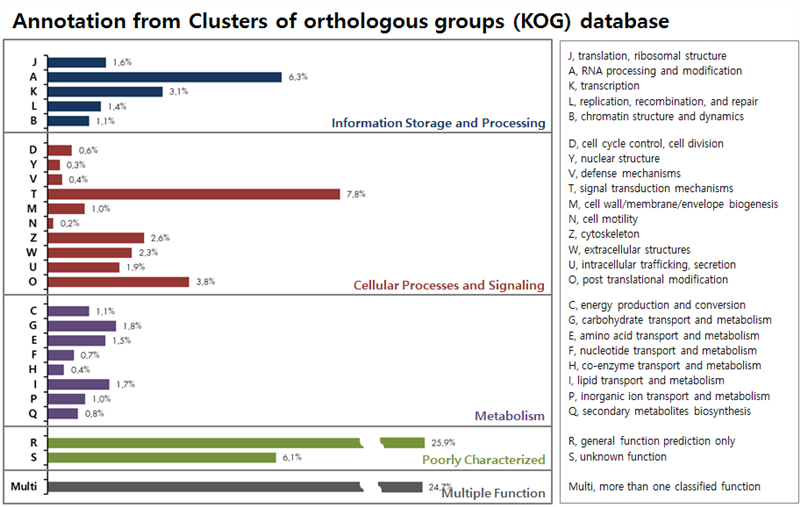
ЛъБМЖвГЊКёРЧ KOG АсАњ, ЁАInfomation Storage and ProcessingЁБ КЮКаПЁМДТ RNA processing and modification АќЗУАњ 6.3%ЗЮ АЁРх ИЙРЬ ИХФЊЕЧАэ, ЁАCellular Processes and SignalingЁБ ПЁМДТ signal transduction mechanisms (7.8%), post translational modification (3.8%) ЗЮ ИЙРЬ ИХФЊЕШДй. ЁАMetabolismЁБ КЮКаРК ЦђБе 1% ДыЗЮ ГЗАд ИХФЊЕЧАэ, Multiple Function РК 24.7% РЬДй.
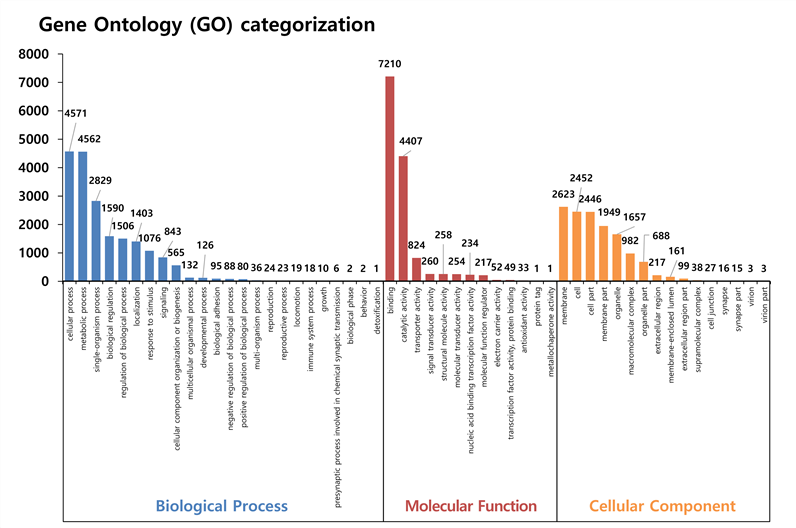
ЛъБМЖвГЊКёРЧ GO АсАњ, Biological ProcessПЁМ cellular process (4,571АГ), metabolic process (4,562АГ) МјРИЗЮ ИЙРЬ ИХФЊЕШДй. Molecular Function ПЁМДТ binding (7,210АГ), catalytic activity (4,407АГ) МјРИЗЮ АЁРх ИЙРЬ ИХФЊЕШДй. ЖЧЧб, Cellular Component КЮКаПЁМДТ membrane (2,623АГ), cell (2,452АГ), cell part (2,446АГ) МјРИЗЮ ИЙРЬ ИХФЊЕШДй.
(4) Microsatellite ШФКИБК
A. SSR statistics
|
ЛъБМЖвГЊКё |
||||||||||
|
Repeats |
4 |
5 |
6 |
7 |
8 |
9 |
10 |
11 |
12 |
13 |
|
Di |
0 |
0 |
300 |
112 |
53 |
34 |
9 |
6 |
5 |
5 |
|
Tri |
0 |
471 |
123 |
41 |
10 |
6 |
1 |
0 |
0 |
0 |
|
Tetra |
596 |
105 |
18 |
9 |
1 |
0 |
0 |
0 |
0 |
0 |
|
Penta |
117 |
38 |
1 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
|
Hexa |
11 |
1 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
0 |
|
Total |
724 |
615 |
442 |
162 |
64 |
40 |
10 |
6 |
5 |
5 |
Ёц SSR АЫЛіАсАњ Уб 2,013АГРЧ SSR МПРЛ ШЎРЮЧв Мі РжОњРИИч, Tetra motif АЁ 729АГЗЮ АЁРх ИЙРЬ СИРчЧЯДТ АЭРЛ ШЎРЮ Чв Мі РжОњРИИч, РЬИІ БтЙнРИЗЮ ЧЯПЉ SSR marker ШФКИБКПЁ ДыЧб primer ИІ 1,436АГ Е№РкРЮ ЧЯПДДй.
B. Primer sequence
Ёц АЂ СОРЧ РЏРќРкИІ КаМЎЧЯПЉ Бз СОРЛ БИКАЧв Мі РжДТ primerИІ ИИЕчДй. primer РЧ СЖАЧРК Motif Ор 4~6АГРЬИч, Forward ПЭ Reverse primer АЁ 18~22АГ СЄЕЕРЬАэ TmАЊРЬ 54.5~55.5 СЄЕЕАЁ РћДчЧЯДй.
- ЛъБМЖвГЊКё
|
Sequence |
Motif |
Forward Primer |
Tm |
|
Reverse Primer |
|||
|
29230117 |
(AGGT)4 |
ACCTACATAACAGTCAAAACGA |
54.14 |
|
AGCCTACTCATCGTCACAGTA |
55.07 |
||
|
29230235 |
(AACA)4 |
TTTTCCGAAATAAAAGGTAGC |
55.48 |
|
TGCATATATTGTGTGCTTTTG |
54.95 |
||
|
29230299 |
(GTAG)4 |
ATATACGTGACAGGAAACACG |
55.22 |
|
GCTATCTTGTGACCGAATAAA |
54.68 |
||
|
29230465 |
(TGAAA)4 |
CAGTGTCATATGTGCCATGTA |
55.39 |
|
CCTGAATAGGAGACAGCAATA |
54.67 |
||
|
29245907 |
(AATA)4 |
GAGATAAGCTGTCCTTGTGC |
55.09 |
|
TCCAGAACTCTGATAGGTACG |
54.56 |
||
|
29246009 |
(GTAC)4 |
GCTACGGAAATAACAATGATG |
54.96 |
|
AGTTTCGAGATCACAGACAGA |
54.98 |
||
|
29246111 |
(TTTA)4 |
GACGTTGGTATATCGTTTGAT |
54.22 |
|
GCGTTGGTAGATCAGTCAGT |
55.3 |
||
|
29246297 |
(TGAC)4 |
ACTAGGGAAAATTGGGTGTAA |
55.36 |
|
TCTAAATCCACAGTTGGTCTG |
55.31 |
||
|
29259255 |
(TTTG)4 |
TTTTCTTTCAACCGAATGAT |
54.86 |
|
ATAATGACCTCGTTATTCTGC |
53.64 |
||
|
29259309 |
(CTGT)4 |
TATTGCAAATGCGAAAGTAAC |
55.74 |
|
CCCTATTTCAACCTCTCAAGT |
55.07 |
||
|
29259339 |
(ATAG)4 |
AACCCCTTTAATAGAGCAAAC |
54.43 |
|
TGATCGTAGCTATAACCCATC |
54.47 |
||
|
29270569 |
(GTTT)4 |
TTATGTACTGGCCTAAATCCA |
54.98 |
|
TTCGCATATATTCCAAGAGTT |
54.23 |
||
|
29270839 |
(AATA)5 |
TGTTTTCATTTGAGCTATGGT |
54.99 |
|
TGAAACGATGACGGTTACTAT |
54.86 |
||
|
29270877 |
(ACACA)4 |
AGTACGCACAGAATCTCTTGA |
55.2 |
|
AGATTTAGCTTCGCATTTTTC |
55.61 |
C. ЙйФкЕх МП
ПфЙј 16ГтЕЕПЁ УЄС§Чб 11СОРЧ РЏРќРкИІ КаМЎЧЯПЉ Бз СОРЛ БИКАЧв Мі РжДТ Barcode МПРЛ ИИЕщОњАэ, Бз Barcode МПРЛ АЁСіАэ NCBI (National Center for Biotechnology Information) ЛчРЬЦЎПЁ РжДТ BLAST (Basic Local Alignment Search Tool)РЛ РЬПыЧЯПЉ АсАњИІ ШЎРЮЧЯПДДй. АЂ СОРЧ МПЕщРЧ BLAST АсАњ ИЖФПЗЮМ СжЗЮ ЛчПыЕЧДТ COI МПАњ ИХФЁЕЧДТ АЭРИЗЮ КИОЦ ЙйФкЕх МПЗЮ ЛчПыАЁДЩЧв АЭРИЗЮ ЛчЗсЕШДй.
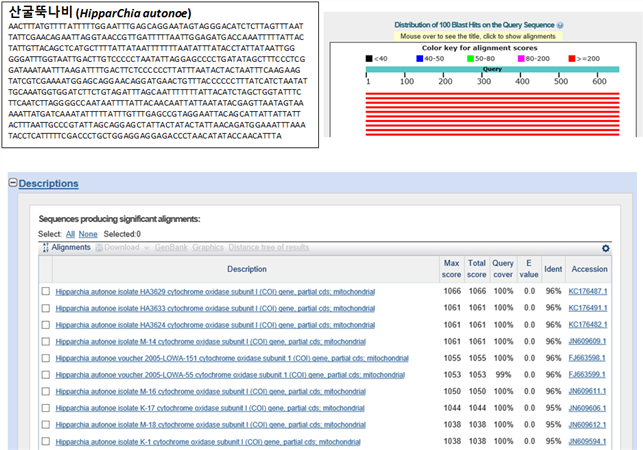
(5) QC АсАњ
|
DNA Library QC |
||||||
|
NO. |
Name |
Concentration (ng/ul) |
Volume (ul) |
Quantity (ug) |
Main peak Size (bp) |
Result |
|
1 |
ЛъБМЖвГЊКё |
87.2 |
3.27 |
285.44 |
470 |
pass |
|
RNA Library QC |
||||||
|
NO. |
Name |
Concentration (ng/ul) |
Volume (ul) |
Quantity (ng) |
Main peak Size (bp) |
Result |
|
1 |
ЛъБМЖвГЊКё |
55.86 |
5.16 |
288.38 |
298 |
Pass |
(6) Gene prediction
|
ЛљЧУСО |
Number of Contig (>1kb) |
Number of Bases (>1kb) |
N50 Contig Size |
Number of gene prediction |
|
ЛъБМЖвГЊКё |
24,308 |
38,803,399 |
1,559 |
12,633 |
АЂСОКАЗЮ transcriptome МПРЛ ДыЛѓРИЗЮ 1kb РЬЛѓИИРЛ МБХУЧЯПЉ Gene predcition Чб АсАњ РЇРЧ ИХФЊЕШ contig, base МіДТ РЇРЧ ЧЅПЭ АААэ N50ЕЧДТ ContigРЧ БцРЬЕЕ КИХы 1~3kb СЄЕЕЗЮ ГЊХИГЕДй. БзИЎАэ РЬЕщРЛ ХыЧи ПЙУјЕШ РЏРќРкРЧ МіДТ 2УЕАГПЁМ 10ИИАГ РЬЛѓРИЗЮ ДйОчЧЯАд ПЙУјЕЧОњДй.
Ёи Long contigРЧ Gene prediction АсАњ БзИВ ПЙНУ