왕은점표범나비 (Fabriciana nerippe)
|
|
▣ 국 명: 왕은점표범나비 ▣ 학 명: Fabriciana nerippe ▣ 지정번호: 멸종위기동물II급 ▣ 계 통: 곤충강-나비목-네발나비과 |
◎ 생김새 : 표범나비류 가운데 중간 크기의 종으로 날개 편 길이는 60~70mm, 앞날개의 길이는 32~44mm이다. 날개 아래쪽에는 은색 무늬의 변이가 다양하게 나타나고 있다. 수컷은 앞날개 윗면에 검은색 줄무늬가 있다. 암컷이 수컷보다 날개 바탕색이 진하며, 날개 아랫면의 은색 무늬도 크고 선명하다. 암컷은 수컷에 비하여 약간 크며 날개의 폭도 약간 넓다. 날개는 주황색으로 검정무늬가 있으며 바깥가두리를 따라 줄지은 무늬의 안쪽은 V자 모양이다. 은점표범나비와 비슷하지만 뒷 날개 뒷면 바깥 가장자리의 산모양의 2개의 은빛 무늬와 수컷 앞날개의 검은색의 1개의 비늘줄로 구별이 가능하다.
◎ 서식지 : 주로 풀밭에서 생활하며, 큰까치수영?엉겅퀴?솔체꽃에 잘 모인다.
◎ 먹이 : 유충은 몇 종의 제비꽃을 먹는다.
◎ 번식 : 1년에 1회 번식하며 따뜻한 곳에서는 5월 하순경부터, 추운 곳에서는 7월 중순경부터 번식힌다.
◎ 분포 : 한국?일본?중국 ?티베트 등지에 분포하며 한반도에는 국지적 분포를 하는 종이다. 내륙 지역에서는 개체밀도가 낮지만 서해안 도서 또는 해안가 지역에서는 자주 관찰된다.
1) 채집정보
- 채집허가: 2014.06.30 / 5마리 , 2014.07.17 / 5마리
- 채 집 자: 빛고을생태연구소 정헌천
2) 채집현황 및 샘플처리
|
왕은점표범나비 채집현황 |
샘플처리 |
|||
|
채집일 |
개체 수 |
DNAseq |
RNAseq |
표본 |
|
2014.07.31 |
5 EA |
2 EA (몸통) |
2 EA (내장) |
1 |
3) QC Data
|
샘플종 |
DNA QC |
RNA QC |
Result |
||
|
Concentration (ug/ul) |
Total Conc. (ug) |
Concentration (ug/ul) |
RIN (>7) |
||
|
왕은점표범나비 |
112.2 |
9 |
430 |
9 |
pass |
4) 기존 NCBI Taxonomy 유전정보 현황
|
유전정보 종류 |
Fabriciana 속 유전자원 |
Fabriciana nerippe 유전자원 |
|
Nucleotide |
342 |
17 |
|
Protein |
364 |
41 |
|
Genome |
1 |
1 |
|
Gene |
13 |
13 |
5) Sequencing 분석 결과
- Illumina DNA Sequncing 결과
|
Input information |
|||||
|
|
|
|
Region1 |
Region2 |
Total |
|
raw data |
|||||
|
Number Of Reads |
|
|
131,791,213 |
131,791,213 |
263,582,426 |
|
Number Of Bases |
|
|
16,605,692,838 |
16,605,692,838 |
33,211,385,676 |
|
error correction |
|||||
|
Number Of Reads |
|
|
129,324,855 |
128,180,861 |
257,505,716 |
|
Number Of Bases |
|
|
15,671,819,041 |
15,486,758,524 |
31,158,577,565 |
|
pair reads |
|||||
|
Number Of Reads |
|
|
|
|
253,514,024 |
|
Number Of Bases |
|
|
|
|
30,721,169,467 |
|
single reads |
|||||
|
Number Of Reads |
|
|
|
|
3,991,692 |
|
Number Of Bases |
|
|
|
|
437,408,098 |
|
Assembly Results |
|||||
|
Scaffold Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Scaffolds |
1,200,140 |
1,200,140 |
123,730 |
35,098 |
7,254 |
|
Number Of Bases |
319,513,086 |
319,513,086 |
118,949,468 |
58,871,980 |
21,680,288 |
|
Avg. Scaffold Size |
266 |
266 |
961 |
1,677 |
2,988 |
|
N50 Scaffold Size |
343 |
343 |
991 |
1,655 |
2,914 |
|
N80 Scaffold Size |
151 |
151 |
638 |
1,195 |
2,268 |
|
N90 Scaffold Size |
113 |
113 |
562 |
1,089 |
2,127 |
|
largest Scaffold Size |
15,105 |
15,105 |
15,105 |
15,105 |
15,105 |
|
Contig Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Contigs |
37,187,522 |
1,659,289 |
42,597 |
6,912 |
887 |
|
Number Of Bases |
1,852,466,098 |
294,652,114 |
33,513,754 |
10,296,573 |
2,455,281 |
|
Avg. Contig Size |
49 |
177 |
786 |
1,489 |
2,768 |
|
N50 Contig Size |
49 |
176 |
755 |
1,432 |
2,616 |
|
N80 Contig Size |
33 |
117 |
570 |
1,126 |
2,193 |
|
N90 Contig Size |
32 |
107 |
531 |
1,057 |
2,085 |
|
Largest Contig Size |
14,052 |
14,052 |
14,052 |
14,052 |
14,052 |
- Illumina RNA Sequncing 결과
|
Input information |
|||||
|
|
Region1 |
Region2 |
Region3 |
Region4 |
Total |
|
raw data |
|||||
|
Number Of Reads |
114,245,854 |
114,245,854 |
8,243,719 |
8,243,719 |
244,979,146 |
|
Number Of Bases |
14,394,977,604 |
14,394,977,604 |
1,038,708,594 |
1,038,708,594 |
30,867,372,396 |
|
error correction |
|||||
|
Number Of Reads |
113,432,927 |
112,934,549 |
7,974,934 |
7,909,590 |
242,252,000 |
|
Number Of Bases |
14,112,009,603 |
14,042,313,816 |
979,327,170 |
970,044,051 |
30,103,694,640 |
|
pair reads |
|||||
|
Number Of Reads |
|
225,107,458 |
|
15,639,746 |
240,747,204 |
|
Number Of Bases |
|
28,016,607,118 |
|
1,924,750,456 |
29,941,357,574 |
|
single reads |
|||||
|
Number Of Reads |
|
1,260,018 |
|
244,778 |
1,504,796 |
|
Number Of Bases |
|
137,716,301 |
|
24,620,765 |
162,337,066 |
|
Assembly Results |
|||||
|
Scaffold Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Scaffolds |
146,801 |
146,801 |
11,040 |
2,633 |
474 |
|
Number Of Bases |
35,154,884 |
35,154,884 |
9,871,862 |
4,231,016 |
1,380,734 |
|
Avg. Scaffold Size |
239 |
239 |
894 |
1,606 |
2,912 |
|
N50 Scaffold Size |
273 |
273 |
891 |
1,558 |
2,765 |
|
N80 Scaffold Size |
144 |
144 |
613 |
1,166 |
2,245 |
|
N90 Scaffold Size |
119 |
119 |
551 |
1,075 |
2,129 |
|
largest Scaffold Size |
9,682 |
9,682 |
9,682 |
9,682 |
9,682 |
|
Contig Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Contigs |
2,159,331 |
181,835 |
7,226 |
1,149 |
103 |
|
Number Of Bases |
161,982,194 |
36,395,092 |
5,643,108 |
1,638,502 |
281,509 |
|
Avg. Contig Size |
75 |
200 |
780 |
1,426 |
2,733 |
|
N50 Contig Size |
78 |
205 |
756 |
1,369 |
2,687 |
|
N80 Contig Size |
51 |
133 |
577 |
1,113 |
2,206 |
|
N90 Contig Size |
47 |
114 |
535 |
1,056 |
2,096 |
|
Largest Contig Size |
5,454 |
5,454 |
5,454 |
5,454 |
5,454 |
(1) 고품질 genomic DNA 채취 및 표본확보
|
검수용 샘플 DNA 농도 및 총 양, 확증표본 현황표 |
|||||||
|
Sample명 |
농도(ng/ul) |
Volume(ul) |
DNA 총량(ug) |
O.D A260/280 |
확증표본 |
채집허가일 |
채집일 |
|
왕은점표범나비 |
388 |
996 |
386 |
1.83 |
○ |
2014.7.17 |
2014.7.31 |
(2) 유전체 서열확보 : 현재 20종 확보
|
Sample |
구분 |
raw data size |
trim data size |
비고 |
|
왕은점표범나비 |
DNA |
33,211,385,676 |
31,158,577,565 |
|
|
RNA |
30,867,372,396 |
30,103,694,640 |
|
(3) 기타분석
A. Genome Size estimation
|
샘 플 명 |
Kmer size |
Kmer total |
Pk depth |
Genome_size |
Coverage(X) |
|
왕은점표범나비 |
23 |
20,559,429,228 |
176 |
116,814,938 |
225.641 |
B. 반복서열
- 시퀀싱이 완료된 왕은점표범나비 DNA 및 RNA의 contigs 중 1kb 이상의 contigs의 반복서열을 탐색(Left; Repeat masking results of DNA contigs, Right; Repeat masking results of RNA contigs).
|
sequences: 6,912 |
|
sequences: 1,240 |
||||||||
|
total length: 10,296,573 bp (10,296,573 bp excl N/X-runs) |
|
total length: 1,779,522bp (1,779,522 bp excl N/X-runs) |
||||||||
|
GC level: 34.62% |
|
GC level: 42.40% |
||||||||
|
bases masked: 4,522 bp (0.04%) |
|
bases masked: 2,763 bp (0.16%) |
||||||||
|
|
number of elements |
length occupied |
percentage of sequence |
|
|
number of elements |
length occupied |
percentage of sequence |
||
|
SINEs: |
0 |
0 bp |
0.00% |
|
SINEs: |
0 |
0 bp |
0.00% |
||
|
|
ALUs |
0 |
0 bp |
0.00% |
|
|
ALUs |
0 |
0 bp |
0.00% |
|
|
MIRs |
0 |
0 bp |
0.00% |
|
|
MIRs |
0 |
0 bp |
0.00% |
|
LINEs: |
4 |
434 bp |
0.00% |
|
LINEs: |
7 |
464 bp |
0.03% |
||
|
|
LINE1 |
1 |
55 bp |
0.00% |
|
|
LINE1 |
0 |
0 bp |
0.00% |
|
|
LINE2 |
0 |
0 bp |
0.00% |
|
|
LINE2 |
1 |
102 bp |
0.01% |
|
|
L3/CR1 |
1 |
38 bp |
0.00% |
|
|
L3/CR1 |
4 |
210 bp |
0.01% |
|
LTR elements: |
4 |
342 bp |
0.00% |
|
LTR elements: |
3 |
426 bp |
0.02% |
||
|
|
ERVL |
1 |
81 bp |
0.00% |
|
|
ERVL |
0 |
0 bp |
0.00% |
|
|
ERVL-MaLRs |
0 |
0 bp |
0.00% |
|
|
ERVL-MaLRs |
0 |
0 bp |
0.00% |
|
|
ERV_classI |
0 |
0 bp |
0.00% |
|
|
ERV_classI |
0 |
0 bp |
0.00% |
|
|
ERV_classII |
0 |
0 bp |
0.00% |
|
|
ERV_classII |
0 |
0 bp |
0.00% |
|
DNA elements: |
14 |
3,221 bp |
0.03% |
|
DNA elements: |
6 |
1,083 bp |
0.06% |
||
|
|
hAT-Charlie |
7 |
1,743 bp |
0.02% |
|
|
hAT-Charlie |
1 |
115 bp |
0.01% |
|
|
TcMar-Tigger |
1 |
90 bp |
0.00% |
|
|
TcMar-Tigger |
0 |
0 bp |
0.00% |
|
Unclassified: |
1 |
41 bp |
0.00% |
|
Unclassified: |
0 |
0 bp |
0.00% |
||
|
Total interspersed repeats: |
|
4,068 bp |
0.04% |
|
Total interspersed repeats: |
|
1,973 bp |
0.11% |
||
|
Small RNA: |
7 |
454 bp |
0.00% |
|
Small RNA: |
3 |
790 bp |
0.04% |
||
|
Satellites: |
0 |
0 bp |
0.00% |
|
Satellites: |
0 |
0 bp |
0.00% |
||
|
Simple repeats: |
0 |
0 bp |
0.00% |
|
Simple repeats: |
0 |
0 bp |
0.00% |
||
|
Low complexity: |
0 |
0 bp |
0.00% |
|
Low complexity: |
0 |
0 bp |
0.00% |
||
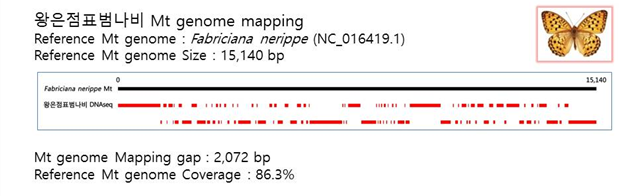
D. Mito-genome Map

(4) Microsatellite 후보군
A. SSR statistics
|
Motif(-mer) |
2 |
3 |
4 |
5 |
6 |
7 |
8 |
9 |
10 |
Total |
|
왕은점표범나비 |
477 |
129 |
4 |
- |
- |
- |
- |
- |
- |
610 |
B. 바코드서열
>왕은점표범나비(Fabriciana nerippe)
TTTAATTTTTCTATCACCCCAACAAAATAATATTAATTAATTTTAATTTATATTTATATA
TTTAATTTTAATTATTAATATTATAAAACTCTATAGGGTCTTCTCGTCTTTTAAATAAAT
TTTAGCTTTTTAACTAAAAAATTAATTTTTATTAATAATTTAGAGACAGTTTATATTTCA
TCAAATCTTTCATACAAGTCTTCAATTAAAAGACTAATGATTATGCTACCTTTGTACAGT
CAATATACTGCAGCCCTTTAATTTTTAATCAGTGGGCAGATTAGACTTTAAATTATTTTC
TAAAAGACATGTTTTTGATAAACAAGTGAATATAAATTTTTGCCGAATTCCTTTAAATTA
ATTTCAATTTATATTTTATATTTTTAATTAATAATATACTAATTTTATCATTATTTCTAT
TTTTTTATTATTAAAAAATATTTTTTTATAAAAAATTAAATAATATTTTAAATTTTATTT
AATATAAAATTTTTTTTAACAAACTATAATTTTTAAACAATATAAATTTTAAAAATTTTA
TTAATATTTATTTTTATTATAAATATTTAAATTAAAGCTTATCCCTTAAAATATTTAATT
TTAAAATTTTATATTTAATTAATTAAATATAAA