대추귀고둥 (Ellobium chinense)
|
|
▣ 국 명: 대추귀고둥 ▣ 학 명: Ellobium chinense ▣ 지정번호: 멸종위기동물 II급 ▣ 계 통: 복족강-진유폐목-대추귀고둥과 |
◎ 생김새 : 껍데기 높이 34mm, 지름 17mm로 껍데기는 대추모양이고 나탑은 원뿔 모양이며 크기가 작은 편이다. 체층이 큰 편이라 껍데기 높이의 4/5에 이른다. 껍데기 주둥이도 커서 껍데기 높이의 약 2/3에 달한다. 껍데기는 갈색의 각피로 덮여 있고 오래된 개체는 흑갈색을 띠며 두꺼운 편이다. 굵고 거친 성장맥이 체층과 다른 층에도 세로로 나있다. 각정에서 체층 둘레의 가장자리까지 성장맥과 나상맥이 교차하며 둘레의 가장자리 아래에서 각저까지는 나상맥이 없고 성장맥만 보이는 특징이 있다. 각정은 둥글고 봉합은 얕다. 체층은 커서 각고의 대부분을 차지고 있으며 각구는 긴 난형으로 외순 아래와 저순은 두껍고 축순에는 강한 주름이 3개 나타난다. 껍데기 주둥이는 좁고 아래위로 긴 반면, 항문이 있는 쪽은 좁고 앞쪽이 둥글며 넓다. 주둥이의 안쪽 입술은 흰색이고 축순에 이빨 모양의 돌기가 1개 있다.
◎ 서식지 : 담수가 들어오는 만조선 부근의 갯벌에서 서식한다.
◎ 연구의 필요성 : 분포지가 좁고 개체수가 줄어들고 있다.
◎ 분포 : 한국의 서해안에 분포한다. 최근 전북 부안?고창의 갯벌 습지 보호지역에서 대규모 서식지가
1) 채집정보
- 채집허가: 2014.07.10 / 2마리
- 채 집 자: 강원대학교 이준상
2) 채집현황 및 샘플처리
|
대추귀고둥 채집현황 |
샘플처리 |
|||
|
채집일 |
개체 수 |
DNAseq |
RNAseq |
표본 |
|
2014.07.19 |
2 EA |
1 EA (발) |
1 EA (내장) |
1 EA, 70% EtOH |
3) QC Data
|
샘플종 |
DNA QC |
RNA QC |
Result |
||
|
Concentration (ug/ul) |
Total Conc. (ug) |
Concentration (ug/ul) |
RIN (>7) |
||
|
대추귀고둥 |
89 |
8.9 |
217 |
8.1 |
pass |
4) 기존 NCBI Taxonomy 유전정보 현황
|
유전정보 종류 |
Ellobium 속 유전자원 |
Ellobium chinense 유전자원 |
|
Nucleotide |
71 |
55 |
|
Protein |
58 |
53 |
|
Genome |
1 |
1 |
|
Gene |
36 |
36 |
5) Sequencing 분석 결과
- Illumina DNA Sequncing 결과
|
Input information |
|||||
|
|
|
|
Region1 |
Region2 |
Total |
|
raw data |
|||||
|
Number Of Reads |
|
|
162,495,135 |
162,495,135 |
324,990,270 |
|
Number Of Bases |
|
|
20,474,387,010 |
20,474,387,010 |
40,948,774,020 |
|
error correction |
|||||
|
Number Of Reads |
|
|
159,557,247 |
158,218,822 |
317,776,069 |
|
Number Of Bases |
|
|
19,066,663,522 |
18,808,352,212 |
37,875,015,734 |
|
pair reads |
|||||
|
Number Of Reads |
|
|
|
|
312,254,234 |
|
Number Of Bases |
|
|
|
|
37,301,833,970 |
|
single reads |
|||||
|
Number Of Reads |
|
|
|
|
5,521,835 |
|
Number Of Bases |
|
|
|
|
573,181,764 |
|
Assembly Results |
|||||
|
Scaffold Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Scaffolds |
1,439,070 |
1,439,070 |
339,344 |
172,177 |
70,402 |
|
Number Of Bases |
745,465,349 |
745,465,349 |
533,106,500 |
415,436,352 |
273,549,102 |
|
Avg. Scaffold Size |
518 |
518 |
1,570 |
2,412 |
3,885 |
|
N50 Scaffold Size |
1,243 |
1,243 |
2,069 |
2,752 |
4,027 |
|
N80 Scaffold Size |
313 |
313 |
944 |
1,513 |
2,605 |
|
N90 Scaffold Size |
167 |
167 |
701 |
1,235 |
2,278 |
|
largest Scaffold Size |
57,718 |
57,718 |
57,718 |
57,718 |
57,718 |
|
Contig Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Contigs |
28,474,636 |
3,007,112 |
258,968 |
57,742 |
8,876 |
|
Number Of Bases |
1,909,388,162 |
728,864,744 |
224,348,524 |
89,196,096 |
24,896,163 |
|
Avg. Contig Size |
67 |
242 |
866 |
1,544 |
2,804 |
|
N50 Contig Size |
68 |
294 |
860 |
1,499 |
2,683 |
|
N80 Contig Size |
37 |
140 |
606 |
1,147 |
2,214 |
|
N90 Contig Size |
33 |
116 |
548 |
1,067 |
2,095 |
|
Largest Contig Size |
20,791 |
20,791 |
20,791 |
20,791 |
20,791 |
- Illumina RNA Sequncing 결과
|
Input information |
|||||
|
|
|
|
Region1 |
Region2 |
Total |
|
raw data |
|||||
|
Number Of Reads |
|
|
176,364,760 |
176,364,760 |
352,729,520 |
|
Number Of Bases |
|
|
22,221,959,760 |
22,221,959,760 |
44,443,919,520 |
|
error correction |
|||||
|
Number Of Reads |
|
|
174,563,243 |
173,659,337 |
348,222,580 |
|
Number Of Bases |
|
|
21,671,483,873 |
21,539,553,316 |
43,211,037,189 |
|
pair reads |
|||||
|
Number Of Reads |
|
|
|
|
345,939,822 |
|
Number Of Bases |
|
|
|
|
42,962,620,927 |
|
single reads |
|||||
|
Number Of Reads |
|
|
|
|
2,282,758 |
|
Number Of Bases |
|
|
|
|
248,416,262 |
|
Assembly Results |
|||||
|
Scaffold Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Scaffolds |
276,425 |
276,425 |
43,438 |
16,772 |
5,207 |
|
Number Of Bases |
97,411,093 |
97,411,093 |
51,367,430 |
32,994,673 |
17,102,164 |
|
Avg. Scaffold Size |
352 |
352 |
1,182 |
1,967 |
3,284 |
|
N50 Scaffold Size |
550 |
550 |
1,355 |
2,061 |
3,239 |
|
N80 Scaffold Size |
195 |
195 |
731 |
1,314 |
2,393 |
|
N90 Scaffold Size |
136 |
136 |
602 |
1,146 |
2,177 |
|
largest Scaffold Size |
21,046 |
21,046 |
21,046 |
21,046 |
21,046 |
|
Contig Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Contigs |
5,047,910 |
395,995 |
33,414 |
8,260 |
1,351 |
|
Number Of Bases |
312,839,041 |
96,828,979 |
29,937,370 |
12,977,658 |
3,814,708 |
|
Avg. Contig Size |
61 |
244 |
895 |
1,571 |
2,823 |
|
N50 Contig Size |
63 |
289 |
902 |
1,527 |
2,680 |
|
N80 Contig Size |
37 |
143 |
617 |
1,161 |
2,221 |
|
N90 Contig Size |
33 |
117 |
553 |
1,074 |
2,110 |
|
Largest Contig Size |
16,764 |
16,764 |
16,764 |
16,764 |
16,764 |
나) 멸종위기종 동물 Genome Survey
|
검수용 샘플 DNA 농도 및 총 양, 확증표본 현황표 |
|||||||
|
Sample명 |
농도(ng/ul) |
Volume(ul) |
DNA 총량(ug) |
O.D A260/280 |
확증표본 |
채집허가일 |
채집일 |
|
대추귀고둥 |
392 |
996 |
390 |
1.83 |
○ |
2014.7.10 |
2014.7.19 |
(2) 유전체 서열확보 : 현재 20종 확보
|
Sample |
구분 |
raw data size |
trim data size |
비고 |
|
대추귀고둥 |
DNA |
40,948,774,020 |
37,875,015,734 |
|
|
RNA |
44,443,919,520 |
43,211,037,189 |
|
(3) 기타분석
A. Genome Size estimation
|
샘 플 명 |
Kmer size |
Kmer total |
Pk depth |
Genome_size |
Coverage(X) |
|
대추귀고둥 |
23 |
25,349,241,060 |
163 |
155,516,816 |
208.974 |
B. 반복서열
- 시퀀싱이 완료된 대추귀고둥 DNA 및 RNA의 contigs 중 1kb 이상의 contigs의 반복서열을 탐색(Left; Repeat masking results of DNA contigs, Right; Repeat masking results of RNA contigs).
|
sequences: 57,742 |
|
sequences: 8,260 |
||||||||
|
totallength:89,196,096bp (89,196,096 bp excl N/X-runs) |
|
totallength:12,977,658bp (12,977,658 bp excl N/X-runs) |
||||||||
|
GC level: 38.74 % |
|
GC level: 43.57 % |
||||||||
|
bases masked: 22,231 bp ( 0.02 %) |
|
bases masked: 3,972 bp ( 0.03 %) |
||||||||
|
|
numberof elements |
length occupied |
percentage of sequence |
|
|
numberof elements |
length occupied |
percentage of sequence |
||
|
SINEs: |
70 |
5,749 bp |
0.01% |
|
SINEs: |
6 |
389 bp |
0.00% |
||
|
|
ALUs |
0 |
0 bp |
0.00% |
|
|
ALUs |
0 |
0 bp |
0.00% |
|
|
MIRs |
39 |
3,652 bp |
0.00% |
|
|
MIRs |
4 |
303 bp |
0.00% |
|
LINEs: |
83 |
6,670 bp |
0.01% |
|
LINEs: |
20 |
1,481 bp |
0.01% |
||
|
|
LINE1 |
10 |
582 bp |
0.00% |
|
|
LINE1 |
6 |
572 bp |
0.00% |
|
|
LINE2 |
8 |
629 bp |
0.00% |
|
|
LINE2 |
3 |
240 bp |
0.00% |
|
|
L3/CR1 |
34 |
1,737 bp |
0.00% |
|
|
L3/CR1 |
4 |
188 bp |
0.00% |
|
LTRelements: |
26 |
1,886 bp |
0.00% |
|
LTRelements: |
6 |
799 bp |
0.01% |
||
|
|
ERVL |
3 |
173 bp |
0.00% |
|
|
ERVL |
0 |
0 bp |
0.00% |
|
|
ERVL-MaLRs |
2 |
124 bp |
0.00% |
|
|
ERVL-MaLRs |
0 |
0 bp |
0.00% |
|
|
ERV_classI |
14 |
873 bp |
0.00% |
|
|
ERV_classI |
4 |
275 bp |
0.00% |
|
|
ERV_classII |
3 |
192 bp |
0.00% |
|
|
ERV_classII |
0 |
0 bp |
0.00% |
|
DNAelements: |
73 |
5,013 bp |
0.01% |
|
DNAelements: |
18 |
1,132 bp |
0.01% |
||
|
|
hAT-Charlie |
15 |
756 bp |
0.02% |
|
|
hAT-Charlie |
5 |
224 bp |
0.00% |
|
|
TcMar-Tigger |
20 |
1,526 bp |
0.00% |
|
|
TcMar-Tigger |
1 |
42 bp |
0.00% |
|
Unclassified: |
4 |
183 bp |
0.00% |
|
Unclassified: |
0 |
0 bp |
0.00% |
||
|
Total interspersed repeats: |
|
19,501 bp |
0.02% |
|
Total interspersed repeats: |
|
3,801 bp |
0.03% |
||
|
Small RNA: |
52 |
2,836 bp |
0.00% |
|
Small RNA: |
3 |
217 bp |
0.00% |
||
|
Satellites: |
0 |
0 bp |
0.00% |
|
Satellites: |
0 |
0 bp |
0.00% |
||
|
Simple repeats: |
0 |
0 bp |
0.00% |
|
Simple repeats: |
0 |
0 bp |
0.00% |
||
|
Low complexity: |
0 |
0 bp |
0.00% |
|
Low complexity: |
0 |
0 bp |
0.00% |
||
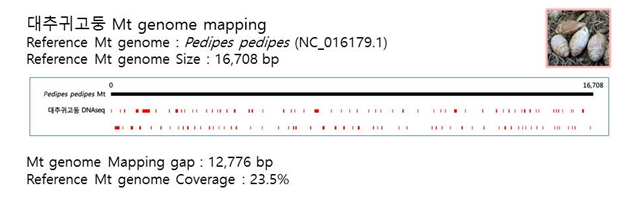
D. Mito-genome Map

(4) Microsatellite 후보군
A. SSR statistics
|
Motif(-mer) |
2 |
3 |
4 |
5 |
6 |
7 |
8 |
9 |
10 |
Total |
|
대추귀고둥 |
8,588 |
3,121 |
1,405 |
124 |
3 |
- |
- |
- |
- |
13,241 |
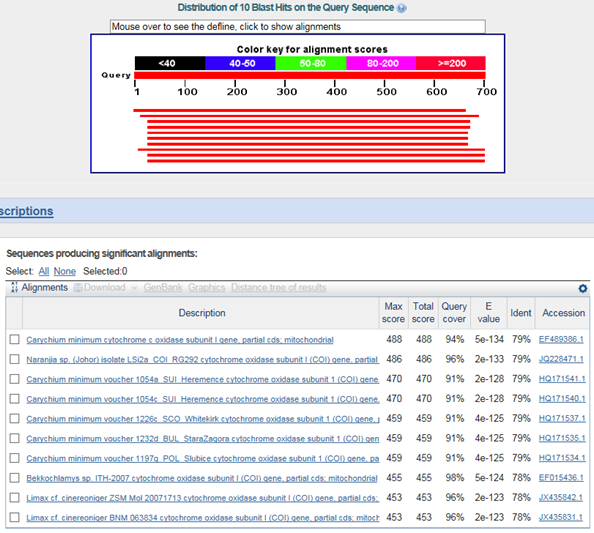
B. 바코드서열
>대추귀고둥(Ellobium chinense)
TCAACAAATCATAAAGATATTGGGACGTTATATATAATTTTTGGGGTTTGGTGTGGCTTA
GTTGGAACTGGGCTGTCACTTCTTATTCGGTTTGAATTAGGAACCGCTGGCAACTTGCTA
GATGATCACTTTTATAATGTTATTGTGACTGCGCACGCTTTTGTGATAATTTTTTTTATG
GTAATACCCTTAATGATCGGGGGGTTCGGAAATTGAATAATTCCTTTGCTTATTGGTGCC
CCGGACATAAGCTTTCCGCGGATAAATAACATGAGTTTTTNNNNNNNNNCTCCATCCTTT
GTTTTATTAATTTTTTCCAGAATAGTTNNNNNNNNNGCTGGGACTGGGTGAACTGTCTAT
CCTCCCTTGAGGGGACCAATTGCTCATGCGGGGGCCTCGGTAGATCTTGTTATTTTCTCG
TTGCATCTTGCCGGAATGTCTTCTATTTTAGGGGCTATTAATTTTATTACTACTGTGTTT
AATATGCGATCGCCGGGAATGACAATGGAACGAGTCAACCTTTTTGTTTGGTCTGTTCTC
GTAACAGTTTTTTTACTTTTACTATCCCTTCCTGTTTTGGCAGGGGCCATTACGATGCTC
CTGACGGATCGAAATTTTAATACTAGTTTTTTTGATCCTGCTGGTGGTGGGGATCCTATT
TTATATCAGCACCTTTTCTGATTTTTTGGTCACCCGGAAGT
Carychium minimum : family 까지 같음
Naranjia sp. (Johor) : superfamily부터 다름