귀이빨대칭이 (Cristaria plicata)
|
|
▣ 국 명: 귀이빨대칭이 ▣ 학 명: Cristaria plicata ▣ 지정번호: 멸종위기야생동물 Ⅰ급 ▣ 계 통: 석패 목, 석패 과 |
1) 채집정보
- 채집허가: 2014.07.01 / 2마리
- 채 집 자: 강원대학교 이준상
2) 채집현황 및 샘플처리
|
귀이빨대칭이 채집현황 |
샘플처리 |
|||
|
채집일 |
개체 수 |
DNAseq |
RNAseq |
표본 |
|
2014.10.25 |
2 EA |
1 EA (발) |
1 EA (내장) |
1 EA |
3) QC Data
|
샘플종 |
DNA QC |
RNA QC |
Result |
||
|
Concentration (ug/ul) |
Total Conc. (ug) |
Concentration (ug/ul) |
RIN (>7) |
||
|
귀이빨 대칭이 |
84.1 |
17 |
349 |
7.4 |
pass |
4) 기존 NCBI Taxonomy 유전정보 현황
|
유전정보 종류 |
Cristaria 속 유전자원 |
Cristaria plicata 유전자원 |
|
Nucleotide |
205 |
194 |
|
Protein |
160 |
159 |
|
Genome |
1 |
1 |
|
Gene |
13 |
13 |
5) Sequencing 분석 결과
- Illumina DNA Sequncing 결과
|
Input information |
|||||
|
|
|
|
Region1 |
Region2 |
Total |
|
raw data |
|||||
|
Number Of Reads |
|
|
147,474,719 |
147,474,719 |
294,949,438 |
|
Number Of Bases |
|
|
18,581,814,594 |
18,581,814,594 |
37,163,629,188 |
|
error correction |
|||||
|
Number Of Reads |
|
|
145,087,202 |
142,393,434 |
287,480,636 |
|
Number Of Bases |
|
|
16,980,344,416 |
16,294,617,173 |
33,274,961,589 |
|
pair reads |
|||||
|
Number Of Reads |
|
|
|
|
280,966,602 |
|
Number Of Bases |
|
|
|
|
32,568,619,699 |
|
single reads |
|||||
|
Number Of Reads |
|
|
|
|
6,514,034 |
|
Number Of Bases |
|
|
|
|
706,341,890 |
|
Assembly Results |
|||||
|
Scaffold Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Scaffolds |
2,399,836 |
2,399,836 |
703,393 |
336,419 |
94,446 |
|
Number Of Bases |
1,210,656,925 |
1,210,656,925 |
866,077,170 |
605,162,126 |
269,474,391 |
|
Avg. Scaffold Size |
504 |
504 |
1,231 |
1,798 |
2,853 |
|
N50 Scaffold Size |
999 |
999 |
1,429 |
1,861 |
2,793 |
|
N80 Scaffold Size |
333 |
333 |
817 |
1,288 |
2,260 |
|
N90 Scaffold Size |
175 |
175 |
653 |
1,137 |
2,122 |
|
largest Scaffold Size |
13,098 |
13,098 |
13,098 |
13,098 |
13,098 |
|
Contig Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Contigs |
19,767,574 |
4,196,594 |
598,163 |
175,313 |
20,811 |
|
Number Of Bases |
2,183,645,096 |
1,285,251,032 |
550,888,568 |
256,864,390 |
52,553,165 |
|
Avg. Contig Size |
110 |
306 |
920 |
1,465 |
2,525 |
|
N50 Contig Size |
157 |
397 |
955 |
1,436 |
2,444 |
|
N80 Contig Size |
53 |
179 |
653 |
1,141 |
2,139 |
|
N90 Contig Size |
47 |
139 |
572 |
1,066 |
2,065 |
|
Largest Contig Size |
11,456 |
11,456 |
11,456 |
11,456 |
11,456 |
- Illumina RNA Sequncing 결과
|
Input information |
|||||
|
|
|
|
Region1 |
Region2 |
Total |
|
raw data |
|||||
|
Number Of Reads |
|
|
143,076,292 |
143,076,292 |
286,152,584 |
|
Number Of Bases |
|
|
18,027,612,792 |
18,027,612,792 |
36,055,225,584 |
|
error correction |
|||||
|
Number Of Reads |
|
|
140,951,928 |
139,872,121 |
280,824,049 |
|
Number Of Bases |
|
|
17,362,946,805 |
17,221,984,593 |
34,584,931,398 |
|
pair reads |
|||||
|
Number Of Reads |
|
|
|
|
278,287,810 |
|
Number Of Bases |
|
|
|
|
34,323,357,010 |
|
single reads |
|||||
|
Number Of Reads |
|
|
|
|
2,536,239 |
|
Number Of Bases |
|
|
|
|
261,574,388 |
|
Assembly Results |
|||||
|
Scaffold Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Scaffolds |
248,154 |
248,154 |
56,916 |
27,544 |
10,956 |
|
Number Of Bases |
122,240,873 |
122,240,873 |
82,066,573 |
61,580,866 |
38,432,323 |
|
Avg. Scaffold Size |
492 |
492 |
1,441 |
2,235 |
3,507 |
|
N50 Scaffold Size |
1,014 |
1,014 |
1,854 |
2,485 |
3,588 |
|
N80 Scaffold Size |
276 |
276 |
877 |
1,458 |
2,488 |
|
N90 Scaffold Size |
171 |
171 |
666 |
1,213 |
2,230 |
|
largest Scaffold Size |
26,930 |
26,930 |
26,930 |
26,930 |
26,930 |
|
Contig Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Contigs |
2,484,327 |
364,285 |
56,402 |
22,173 |
6,234 |
|
Number Of Bases |
262,366,772 |
126,715,567 |
64,270,057 |
40,620,088 |
18,731,695 |
|
Avg. Contig Size |
105 |
347 |
1,139 |
1,831 |
3,004 |
|
N50 Contig Size |
91 |
513 |
1,292 |
1,895 |
2,934 |
|
N80 Contig Size |
57 |
189 |
726 |
1,277 |
2,301 |
|
N90 Contig Size |
47 |
146 |
601 |
1,131 |
2,144 |
|
Largest Contig Size |
15,737 |
15,737 |
15,737 |
15,737 |
15,737 |
(1) 고품질 genomic DNA 채취 및 표본확보
|
검수용 샘플 DNA 농도 및 총 양, 확증표본 현황표 |
|||||||
|
Sample명 |
농도(ng/ul) |
Volume(ul) |
DNA 총량(ug) |
O.D A260/280 |
확증표본 |
채집허가일 |
채집일 |
|
귀이빨대칭이 |
602 |
996 |
599 |
2.08 |
○ |
2014.7.1 |
2014.10.25 |
(2) 유전체 서열확보 : 현재 20종 확보
|
Sample |
구분 |
raw data size |
trim data size |
비고 |
|
귀이빨대칭이 |
DNA |
37,163,629,188 |
33,274,961,589 |
|
|
RNA |
36,055,225,584 |
34,584,931,398 |
|
(3) 기타분석
A. Genome Size estimation
|
샘 플 명 |
Kmer size |
Kmer total |
Pk depth |
Genome_size |
Coverage(X) |
|
귀이빨대칭이 |
23 |
23,006,056,164 |
7 |
3,286,579,452 |
8.97436 |
B. 반복서열
- 시퀀싱이 완료된 귀이빨대칭이 DNA 및 RNA의 contigs 중 1kb 이상의 contigs의 반복서열을 탐색(Left; Repeat masking results of DNA contigs, Right; Repeat masking results of RNA contigs).
|
sequences: 175,313 |
|
sequences: 22,173 |
||||||||
|
total length: 256,864,390 bp (256,864,390 bp excl N/X-runs) |
|
total length: 40,620,088bp (40,620,088 bp excl N/X-runs) |
||||||||
|
GC level: 34.58% |
|
GC level: 37.09% |
||||||||
|
bases masked: 1,027,933 bp (0.40%) |
|
bases masked: 61,437 bp (0.15%) |
||||||||
|
|
number of elements |
length occupied |
percentage of sequence |
|
|
number of elements |
length occupied |
percentage of sequence |
||
|
SINEs: |
213 |
12,085 |
0.00% |
|
SINEs: |
26 |
1,587 bp |
0.00% |
||
|
|
ALUs |
0 |
0 |
0.00% |
|
|
ALUs |
0 |
0 bp |
0.00% |
|
|
MIRs |
76 |
4,930 |
0.00% |
|
|
MIRs |
10 |
681 bp |
0.00% |
|
LINEs: |
3,402 |
878,128 |
0.34% |
|
LINEs: |
158 |
38,248 bp |
0.09% |
||
|
|
LINE1 |
81 |
5,526 |
0.00% |
|
|
LINE1 |
12 |
761 bp |
0.00% |
|
|
LINE2 |
483 |
49,882 |
0.02% |
|
|
LINE2 |
19 |
1,962 bp |
0.00% |
|
|
L3/CR1 |
148 |
10,775 |
0.00% |
|
|
L3/CR1 |
12 |
1,086 bp |
0.00% |
|
LTR elements: |
136 |
16,225 |
0.01% |
|
LTR elements: |
25 |
2,373 bp |
0.01% |
||
|
|
ERVL |
7 |
433 |
0.00% |
|
|
ERVL |
2 |
112 bp |
0.00% |
|
|
ERVL-MaLRs |
3 |
133 |
0.00% |
|
|
ERVL-MaLRs |
1 |
64 bp |
0.00% |
|
|
ERV_classI |
103 |
11,930 |
0.00% |
|
|
ERV_classI |
15 |
986 bp |
0.00% |
|
|
ERV_classII |
7 |
435 |
0.00% |
|
|
ERV_classII |
4 |
327 bp |
0.00% |
|
DNA elements: |
1,007 |
86,630 |
0.03% |
|
DNA elements: |
138 |
14,663 bp |
0.04% |
||
|
|
hAT-Charlie |
204 |
25,063 |
0.01% |
|
|
hAT-Charlie |
37 |
5,773 bp |
0.01% |
|
|
TcMar-Tigger |
192 |
19,924 |
0.01% |
|
|
TcMar-Tigger |
32 |
4,005 bp |
0.01% |
|
Unclassified: |
11 |
616 |
0.00% |
|
Unclassified: |
2 |
128 bp |
0.00% |
||
|
Total interspersed repeats: |
|
993,684 |
0.39% |
|
Total interspersed repeats: |
|
56,999 bp |
0.14% |
||
|
Small RNA: |
635 |
37,708 |
0.01% |
|
Small RNA: |
50 |
4,603 bp |
0.01% |
||
|
Satellites: |
10 |
1,311 |
0.00% |
|
Satellites: |
1 |
167 bp |
0.00% |
||
|
Simple repeats: |
0 |
0 |
0.00% |
|
Simple repeats: |
0 |
0 bp |
0.00% |
||
|
Low complexity: |
0 |
0 |
0.00% |
|
Low complexity: |
0 |
0 bp |
0.00% |
||
D. Mito-genome Map
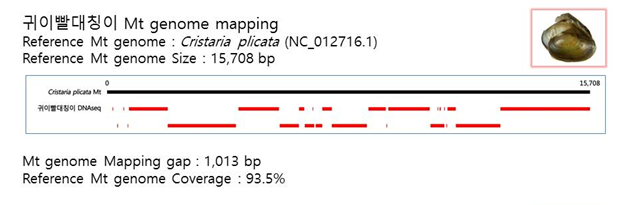
(4) Microsatellite 후보군
A. SSR statistics
|
Motif(-mer) |
2 |
3 |
4 |
5 |
6 |
7 |
8 |
9 |
10 |
Total |
|
귀이빨대칭이 |
27,882 |
7,001 |
5,073 |
140 |
6 |
- |
- |
1 |
1 |
40,104 |
B. 바코드서열
>귀이빨대칭이(Cristaria plicata)
CCGGTCTTAACTCAGCTCGTGTAAGGGTATAATAGTCGAACAGACTATCCTGTCATAGCG
GTTGCACTAATGTGGATATCCTTAAGCCAACATCGAGGTCGCAAACCCAACTTTCGATAT
GTACTCTTGAGTTGGATTACGCTGTTATCCCCGGGGTAGCTTCTTTTTGGTCCTTAGTAA
GTTTTAGATGTGATTTCTAAAGAATCGGAAGCTTTGATGATTCCGAGATTGCCCCAATCA
AACCTTAAGCACTTTTAAGTGTGTAAACAGAAGCGCGAGAAGAGGTAAAGCTCCGCGGGG
TCTTTTCGTCTTCCTTTGTTATCTAAGTCTTTTCACTTAAACGAAAAGTTTTTTTGACAC
CACAAAGTAAAGGTGTTCCTAGTCTTGCCATTCACTGGCCTCCAATTAAAAGGCTAATTA
TTACGCTACCTTAGCACGCTCACGCTAACGCGGCCGTTTAAAGTTTCACTGGGCAGGCAT
TATCTTTTATCAGAGACTTCAAAAGACGATGTTTTTGGTAAACAGGC