기수갈고둥 (Clithon retropictus)
|
|
▣ 국 명: 기수갈고둥 ▣ 학 명: Clithon retropictus ▣ 지정번호: 멸종위기야생동물 Ⅱ급 ▣ 계 통: 복족강-원시복족목-갈고둥과 |
◎ 생김새 : 껍데기 높이 13.5mm, 지름 12.4mm로 전체적으로 작고 겉보기에 공처럼 생겼으며 염주알다슬기와 비슷하다. 껍데기는 두껍고 단단한 편이며 체층이 커서 거의 전부를 차지하고 있어 상대적으로 나탑이 작고 낮은 특징을 보인다. 껍데기의 주둥이는 반달 모양이며 안쪽 입술은 곧으며 가운데 나와 있는 몇 개의 이빨처럼 생긴 돌기중 한 개가 특히 발달하고 있다. 활층은 넓고 편평하다. 껍데기는 갈색의 각피로 덮여 있고 작은 삼각형 모양의 노란빛을 띤 검은색 반점이 있다. 뚜껑은 반원 모양이고 석회질로 이루어져 있으며 주둥이를 덮어 막을 수 있다. 뚜껑의 표면은 매끌매끌하지만 안쪽 가장자리는 얕게 패어있고, 내면에는 석회질의 돌기가 길게 나 있다.
◎ 서식지 : 물의 속도가 빠르고 모래와 자갈이 섞인 하천의 기수지역에 서식하면서 담수 또는 하천의 하구 가까이에도 서식한다.
◎ 번식 : 7~8월 무렵 타원형의 알주머니를 바위나 조개껍데기 위에 붙인다.
◎ 연구의 필요성 : 분포지가 좁고 개체수가 줄어들고 있다.
◎ 분포 : 한국의 남해안과 일본 등지에 분포한다. 최근 경북 울진군 왕피천 하류에서 집단 서식이 확인 되었다.
1) 채집정보
- 채집허가: 2014.07.10 / 5마리
- 채 집 자: 강원대학교 이준상
2) 채집현황 및 샘플처리
|
기수갈고둥 채집현황 |
샘플처리 |
|||
|
채집일 |
개체 수 |
DNAseq |
RNAseq |
표본 |
|
2014.07.19 |
5 EA |
4 EA (발) |
4 EA (내장) |
1 EA, 70% EtOH |
3) QC Data
|
샘플종 |
DNA QC |
RNA QC |
Result |
||
|
Concentration (ug/ul) |
Total Conc. (ug) |
Concentration (ug/ul) |
RIN (>7) |
||
|
기수갈고둥 |
98 |
7.8 |
285 |
9.5 |
pass |
4) 기존 NCBI Taxonomy 유전정보 현황
|
유전정보 종류 |
Clithon 속 유전자원 |
Clithon retropictus 유전자원 |
|
Nucleotide |
191 |
79 |
|
Protein |
143 |
52 |
|
Genome |
1 |
1 |
|
Gene |
74 |
74 |
5) Sequencing 분석 결과
- Illumina DNA Sequncing 결과
|
Input information |
|||||
|
|
|
|
Region1 |
Region2 |
Total |
|
raw data |
|||||
|
Number Of Reads |
|
|
120,273,976 |
120,273,976 |
240,547,952 |
|
Number Of Bases |
|
|
15,154,520,976 |
15,154,520,976 |
30,309,041,952 |
|
error correction |
|||||
|
Number Of Reads |
|
|
117,834,355 |
116,458,835 |
234,293,190 |
|
Number Of Bases |
|
|
14,156,727,932 |
13,863,352,950 |
28,020,080,882 |
|
pair reads |
|||||
|
Number Of Reads |
|
|
|
|
229,934,032 |
|
Number Of Bases |
|
|
|
|
27,573,617,762 |
|
single reads |
|||||
|
Number Of Reads |
|
|
|
|
4,359,158 |
|
Number Of Bases |
|
|
|
|
446,463,120 |
|
Assembly Results |
|||||
|
Scaffold Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Scaffolds |
1,216,526 |
1,216,526 |
368,204 |
182,280 |
67,257 |
|
Number Of Bases |
695,767,098 |
695,767,098 |
518,139,249 |
387,154,839 |
227,044,748 |
|
Avg. Scaffold Size |
571 |
571 |
1,407 |
2,123 |
3,375 |
|
N50 Scaffold Size |
1,192 |
1,192 |
1,745 |
2,312 |
3,392 |
|
N80 Scaffold Size |
386 |
386 |
878 |
1,407 |
2,436 |
|
N90 Scaffold Size |
206 |
206 |
674 |
1,189 |
2,206 |
|
largest Scaffold Size |
22,123 |
22,123 |
22,123 |
22,123 |
22,123 |
|
Contig Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Contigs |
26,095,075 |
2,296,410 |
326,840 |
96,655 |
17,382 |
|
Number Of Bases |
1,687,416,521 |
695,210,658 |
311,508,392 |
153,851,908 |
47,866,494 |
|
Avg. Contig Size |
64 |
302 |
953 |
1,591 |
2,753 |
|
N50 Contig Size |
64 |
431 |
990 |
1,575 |
2,653 |
|
N80 Contig Size |
34 |
171 |
647 |
1,175 |
2,207 |
|
N90 Contig Size |
32 |
126 |
568 |
1,081 |
2,098 |
|
Largest Contig Size |
11,073 |
11,073 |
11,073 |
11,073 |
11,073 |
- Illumina RNA Sequncing 결과
|
Input information |
|||||
|
|
|
|
Region1 |
Region2 |
Total |
|
raw data |
|||||
|
Number Of Reads |
|
|
123,371,899 |
123,371,899 |
246,743,798 |
|
Number Of Bases |
|
|
15,544,859,274 |
15,544,859,274 |
31,089,718,548 |
|
error correction |
|||||
|
Number Of Reads |
|
|
121,345,743 |
120,351,070 |
241,696,813 |
|
Number Of Bases |
|
|
15,021,994,342 |
14,868,772,951 |
29,890,767,293 |
|
pair reads |
|||||
|
Number Of Reads |
|
|
|
|
239,293,972 |
|
Number Of Bases |
|
|
|
|
29,634,758,626 |
|
single reads |
|||||
|
Number Of Reads |
|
|
|
|
2,402,841 |
|
Number Of Bases |
|
|
|
|
256,008,667 |
|
Assembly Results |
|||||
|
Scaffold Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Scaffolds |
254,806 |
254,806 |
43,653 |
17,687 |
5,936 |
|
Number Of Bases |
95,927,319 |
95,927,319 |
53,942,723 |
36,060,688 |
19,910,337 |
|
Avg. Scaffold Size |
376 |
376 |
1,235 |
2,038 |
3,354 |
|
N50 Scaffold Size |
625 |
625 |
1,456 |
2,189 |
3,337 |
|
N80 Scaffold Size |
209 |
209 |
753 |
1,347 |
2,427 |
|
N90 Scaffold Size |
141 |
141 |
611 |
1,160 |
2,199 |
|
largest Scaffold Size |
16,441 |
16,441 |
16,441 |
16,441 |
16,441 |
|
Contig Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Contigs |
4,152,745 |
369,467 |
36,220 |
9,859 |
1,692 |
|
Number Of Bases |
263,748,027 |
95,536,735 |
33,508,299 |
15,600,852 |
4,690,610 |
|
Avg. Contig Size |
63 |
258 |
925 |
1,582 |
2,772 |
|
N50 Contig Size |
63 |
321 |
947 |
1,559 |
2,667 |
|
N80 Contig Size |
36 |
148 |
631 |
1,168 |
2,201 |
|
N90 Contig Size |
33 |
119 |
560 |
1,078 |
2,091 |
|
Largest Contig Size |
9,674 |
9,674 |
9,674 |
9,674 |
9,674 |
나) 멸종위기종 동물 Genome Survey
|
검수용 샘플 DNA 농도 및 총 양, 확증표본 현황표 |
|||||||
|
Sample명 |
농도(ng/ul) |
Volume(ul) |
DNA 총량(ug) |
O.D A260/280 |
확증표본 |
채집허가일 |
채집일 |
|
기수갈고둥 |
309 |
996 |
307 |
1.95 |
○ |
2014.7.10 |
2014.7.19 |
(2) 유전체 서열확보 : 현재 20종 확보
|
Sample |
구분 |
raw data size |
trim data size |
비고 |
|
기수갈고둥 |
DNA |
30,309,041,952 |
28,020,080,882 |
|
|
RNA |
31,089,718,548 |
29,890,767,293 |
|
(3) 기타분석
A. Genome Size estimation
|
샘 플 명 |
Kmer size |
Kmer total |
Pk depth |
Genome_size |
Coverage(X) |
|
기수갈고둥 |
23 |
18,762,740,256 |
155 |
121,049,937 |
198.718 |
B. 반복서열
- 시퀀싱이 완료된 기수갈고둥 DNA 및 RNA의 contigs 중 1kb 이상의 contigs의 반복서열을 탐색(Left; Repeat masking results of DNA contigs, Right; Repeat masking results of RNA contigs).
|
sequences: 96,655 |
|
sequences: 9,859 |
||||||||
|
total length: 153,851,908 bp (153,851,908 bp excl N/X-runs) |
|
total length: 15,600,852 bp (15600852 bp excl N/X-runs) |
||||||||
|
GClevel:44.06% |
|
GC level: 49.17 % |
||||||||
|
bases masked: 69,346 bp ( 0.05 %) |
|
bases masked: 5,524 bp ( 0.04 %) |
||||||||
|
|
numberof elements |
length occupied |
percentage of sequence |
|
|
numberof elements |
length occupied |
percentage of sequence |
||
|
SINEs: |
35 |
2,252 bp |
0.00% |
|
SINEs: |
3 |
137 bp |
0.00% |
||
|
|
ALUs |
0 |
0 bp |
0.00% |
|
|
ALUs |
0 |
0 bp |
0.00% |
|
|
MIRs |
5 |
271 bp |
0.00% |
|
|
MIRs |
1 |
47 bp |
0.00% |
|
LINEs: |
352 |
52,749 bp |
0.03% |
|
LINEs: |
26 |
3,736 bp |
0.02% |
||
|
|
LINE1 |
40 |
2,177 bp |
0.00% |
|
|
LINE1 |
5 |
283 bp |
0.00% |
|
|
LINE2 |
49 |
3,406 bp |
0.00% |
|
|
LINE2 |
7 |
782 bp |
0.01% |
|
|
L3/CR1 |
39 |
2,334 bp |
0.00% |
|
|
L3/CR1 |
4 |
270 bp |
0.00% |
|
LTRelements: |
62 |
4,124 bp |
0.00%. |
|
LTRelements: |
8 |
620 bp |
0.00% |
||
|
|
ERVL |
6 |
458 bp |
0.00% |
|
|
ERVL |
3 |
248 bp |
0.00% |
|
|
ERVL-MaLRs |
2 |
135 bp |
0.00% |
|
|
ERVL-MaLRs |
1 |
37 bp |
0.00% |
|
|
ERV_classI |
34 |
2,109 bp |
0.00% |
|
|
ERV_classI |
1 |
74 bp |
0.00% |
|
|
ERV_classII |
9 |
445 bp |
0.00% |
|
|
ERV_classII |
0 |
0 bp |
0.00% |
|
DNAelements: |
72 |
6,039 bp |
0.00% |
|
DNAelements: |
9 |
961 bp |
0.01% |
||
|
|
hAT-Charlie |
8 |
476 bp |
0.00% |
|
|
hAT-Charlie |
1 |
86 bp |
0.00% |
|
|
TcMar-Tigger |
22 |
2,379 bp |
0.00% |
|
|
TcMar-Tigger |
3 |
215 bp |
0.00% |
|
Unclassified: |
6 |
407 bp |
0.00% |
|
Unclassified: |
0 |
0 bp |
0.00% |
||
|
Total interspersed repeats: |
|
65,571 bp |
0.04% |
|
Total interspersed repeats: |
|
5454 bp |
0.03% |
||
|
Small RNA: |
70 |
3,842 bp |
0.00% |
|
Small RNA: |
1 |
70 bp |
0.00% |
||
|
Satellites: |
1 |
137 bp |
0.00% |
|
Satellites: |
0 |
0 bp |
0.00% |
||
|
Simple repeats: |
0 |
0 bp |
0.00% |
|
Simple repeats: |
0 |
0 bp |
0.00% |
||
|
Low complexity: |
0 |
0 bp |
0.00% |
|
Low complexity: |
0 |
0 bp |
0.00% |
||
D. Mito-genome Map
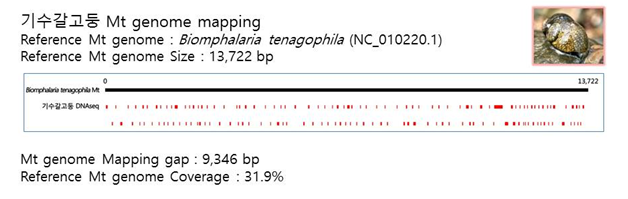
(4) Microsatellite 후보군
A. SSR statistics
|
Motif(-mer) |
2 |
3 |
4 |
5 |
6 |
7 |
8 |
9 |
10 |
Total |
|
기수갈고둥 |
48,716 |
8,728 |
5,666 |
564 |
11 |
- |
- |
- |
- |
63,685 |
B. 바코드서열
>기수갈고둥(Clithon retropictus)
GAGCTTGGACAGCCTGGTGCCTTGCTTGGGGATGATCAATTATATAATGTAATTGTGACT
GCTCATGCGTTTGTGATAATTTTCTTTTTGGTTATACCAATGATGATTGGTGGGTTTGGT
AACTGATTAGTGCCTTTAATGCTAGGGGCTCCTGACATGGCCTTCCCCCGTCTTAATAAC
ATGAGTTTTTGATTACTTCCACCATCATTAACTTTATTATTAGCCTCCTCTGCTGTTGAG
AGTGGAGTAGGGACTGGTTGAACTGTTTATCCTCCATTGTCTGGTAATTTAGCACATGCC
GGTGGGTCGGTTGATTTAGCTATTTTTTCTCTTCACTTAGCTGGTGTTTCTTCAATTTTG
GGTGCAGTAAATTTTATTACCACAATTATTAATATGCGATGGCAGGGAATGCAGTTTGAA
CGATTGCCATTATTTGTTTGATCTGTAAAGATTACTGCAATTTTGCTCTTATTATCACTA
CCTGTGTTGGCAGGAGCAATTACTATATTATTAACTGATCGAAATTTTAATACATCATTT
TTTGATCCAGCAGGTGGGGGTGACCCAATCTTATATCAACA
Clithon spinosus : family 까지 같음
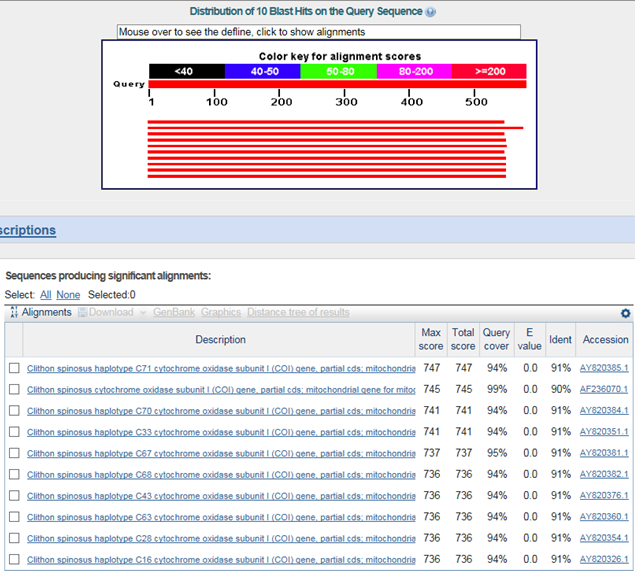 (1) 고품질 genomic DNA 채취 및 표본확보
(1) 고품질 genomic DNA 채취 및 표본확보
