갯게 (Chasmagnathus convexus)
|
|
▣ 국 명: 갯게 ▣ 영 명: mud-flat crab ▣ 학 명: Chasmagnathus convexus ▣ 지정번호: 멸종위기야생동물Ⅱ급 ▣ 계 통: 갑각강-십각목-참게과 |
◎ 생김새 : 갑각의 윤곽은 양 옆가장자리가 볼록한 사각형인데 그 길이는 나비의 10/13정도이다. 이마는 짧은 혀모양이어서 그 가장자리가 둥그스름하고 아래쪽으로 매우 기운 것이 특징이다. 등면에는 깊은 세로홈이 있고 이 홈은 가운데 위구역에까지 뻗는다. 눈구멍 뒷 가장자리는 뚜렷한 두둑을 이룬다. 갑각의 옆가장자리는 매우 볼록하고 눈 뒷니를 포함하여 4개의 넓은 이가 있는데 맨 뒤의 것은 구분이 뚜렷하지 않다.
◎ 서식지 : 하구 가까이의 민물이 흐르는 도랑 또는 갈대밭에 구멍을 파고 살며 상조선 근처에서 깊은 구멍을 파고 사는 것도 있다. 서식지가 도로 건설등에 의하여 파괴되어 멸조위기에 처한 종이다.
◎ 분포 : 한국?일본?지에 분포하며 한국에서는 인천 강화도와 남해도의 갈대밭에 서식한다. 최근 제주도 모슬포와 여수 연안에서도 서식이 확인되고 있다.
1) 채집정보
- 채집허가: 2014.07.10 / 5마리
- 채 집 자: 신라대학교 고현숙
2) 채집현황 및 샘플처리
|
갯게 채집현황 |
샘플처리 |
|||
|
채집일 |
개체 수 |
DNAseq |
RNAseq |
표본 |
|
2014.08.27 |
2 EA |
2 EA (몸통) |
2 EA (내장) |
- |
3) QC Data
|
샘플종 |
DNA QC |
RNA QC |
Result |
||
|
Concentration (ug/ul) |
Total Conc. (ug) |
Concentration (ug/ul) |
RIN (>7) |
||
|
갯게 |
45.7 |
3.6 |
606.6 |
6.7 |
pass |
4) 기존 NCBI Taxonomy 유전정보 현황
|
유전정보 종류 |
Chasmagnathus 속 유전자원 |
Chasmagnathus convexus 유전자원 |
|
Nucleotide |
18 |
18 |
|
Protein |
3 |
3 |
5) Sequencing 분석 결과
- Illumina DNA Sequncing 결과
|
Input information |
|||||
|
|
|
|
Region1 |
Region2 |
Total |
|
raw data |
|||||
|
Number Of Reads |
|
|
153,623,443 |
153,623,443 |
307,246,886 |
|
Number Of Bases |
|
|
19,356,553,818 |
19,356,553,818 |
38,713,107,636 |
|
error correction |
|||||
|
Number Of Reads |
|
|
150,790,723 |
149,270,752 |
300,061,475 |
|
Number Of Bases |
|
|
18,299,853,139 |
18,132,103,317 |
36,431,956,456 |
|
pair reads |
|||||
|
Number Of Reads |
|
|
|
|
295,475,870 |
|
Number Of Bases |
|
|
|
|
35,919,383,637 |
|
single reads |
|||||
|
Number Of Reads |
|
|
|
|
4,585,605 |
|
Number Of Bases |
|
|
|
|
512,572,819 |
|
Assembly Results |
|||||
|
Scaffold Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Scaffolds |
1,861,541 |
1,861,541 |
459,931 |
225,587 |
80,411 |
|
Number Of Bases |
917,173,752 |
917,173,752 |
642,936,252 |
477,829,625 |
275,878,606 |
|
Avg. Scaffold Size |
492 |
492 |
1,397 |
2,118 |
3,430 |
|
N50 Scaffold Size |
1,071 |
1,071 |
1,711 |
2,279 |
3,412 |
|
N80 Scaffold Size |
301 |
301 |
870 |
1,396 |
2,435 |
|
N90 Scaffold Size |
160 |
160 |
671 |
1,185 |
2,203 |
|
largest Scaffold Size |
34,968 |
34,968 |
34,968 |
34,968 |
34,968 |
|
Contig Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Contigs |
20,933,404 |
3,549,408 |
410,643 |
118,957 |
21,539 |
|
Number Of Bases |
1,988,515,949 |
978,693,506 |
391,546,899 |
192,032,899 |
62,192,297 |
|
Avg. Contig Size |
94 |
275 |
953 |
1,614 |
2,887 |
|
N50 Contig Size |
96 |
365 |
985 |
1,580 |
2,768 |
|
N80 Contig Size |
50 |
153 |
645 |
1,176 |
2,231 |
|
N90 Contig Size |
46 |
124 |
567 |
1,082 |
2,110 |
|
Largest Contig Size |
15,357 |
15,357 |
15,357 |
15,357 |
15,357 |
- Illumina RNA Sequncing 결과
|
Input information |
|||||
|
|
|
|
Region1 |
Region2 |
Total |
|
raw data |
|||||
|
Number Of Reads |
|
|
162,451,399 |
162,451,399 |
324,902,798 |
|
Number Of Bases |
|
|
20,468,876,274 |
20,468,876,274 |
40,937,752,548 |
|
error correction |
|||||
|
Number Of Reads |
|
|
160,563,707 |
159,222,631 |
319,786,338 |
|
Number Of Bases |
|
|
19,866,633,265 |
19,690,735,691 |
39,557,368,956 |
|
pair reads |
|||||
|
Number Of Reads |
|
|
|
|
317,265,048 |
|
Number Of Bases |
|
|
|
|
39,282,129,074 |
|
single reads |
|||||
|
Number Of Reads |
|
|
|
|
2,521,290 |
|
Number Of Bases |
|
|
|
|
275,239,882 |
|
Assembly Results |
|||||
|
Scaffold Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Scaffolds |
193,895 |
193,895 |
33,702 |
13,647 |
4,339 |
|
Number Of Bases |
73,081,363 |
73,081,363 |
40,849,327 |
26,962,339 |
14,173,580 |
|
Avg. Scaffold Size |
376 |
376 |
1,212 |
1,975 |
3,266 |
|
N50 Scaffold Size |
619 |
619 |
1,406 |
2,087 |
3,252 |
|
N80 Scaffold Size |
209 |
209 |
753 |
1,324 |
2,392 |
|
N90 Scaffold Size |
145 |
145 |
612 |
1,148 |
2,185 |
|
largest Scaffold Size |
21,944 |
21,944 |
21,944 |
21,944 |
21,944 |
|
Contig Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Contigs |
4,023,890 |
255,107 |
30,210 |
9,990 |
2,178 |
|
Number Of Bases |
236,793,481 |
73,420,982 |
30,690,612 |
16,822,864 |
6,239,197 |
|
Avg. Contig Size |
58 |
287 |
1,015 |
1,683 |
2,864 |
|
N50 Contig Size |
62 |
382 |
1,085 |
1,696 |
2,775 |
|
N80 Contig Size |
34 |
161 |
668 |
1,209 |
2,246 |
|
N90 Contig Size |
32 |
125 |
577 |
1,097 |
2,109 |
|
Largest Contig Size |
11,371 |
11,371 |
11,371 |
11,371 |
11,371 |
나) 멸종위기종 동물 Genome Survey
|
검수용 샘플 DNA 농도 및 총 양, 확증표본 현황표 |
|||||||
|
Sample명 |
농도(ng/ul) |
Volume(ul) |
DNA 총량(ug) |
O.D A260/280 |
확증표본 |
채집허가일 |
채집일 |
|
갯게 (수컷) |
110 |
996 |
109 |
1.56 |
X |
2014.7.10 |
2014.8.27 |
(2) 유전체 서열확보 : 현재 20종 확보
|
Sample |
구분 |
raw data size |
trim data size |
비고 |
|
갯게 |
DNA |
38,713,107,636 |
36,431,956,456 |
|
|
RNA |
40,937,752,548 |
39,557,368,956 |
|
(3) 기타분석
A. Genome Size estimation
|
샘 플 명 |
Kmer size |
Kmer total |
Pk depth |
Genome_size |
Coverage(X) |
|
갯게 |
23 |
23,965,257,108 |
9 |
2,662,806,345 |
11.5385 |
B. 반복서열
- 시퀸싱이 완료된 갯게 DNA 및 RNA의 contigs 중 1kb 이상의 contigs의 반복서열을 탐색(Left; Repeat masking results of DNA contigs, Right; Repeat masking results of RNA contigs).
|
sequences: 118,957 |
|
sequences: 9,990 |
||||||||
|
total length: 192,032,899 bp (192,032,899 bp excl N/X-runs) |
|
total length: 16,822,864bp (16,822,864bp excl N/X-runs) |
||||||||
|
GC level: 42.55% |
|
GC level: 47.17% |
||||||||
|
bases masked: 564,853 bp (0.29%) |
|
bases masked: 21,554 bp (0.13%) |
||||||||
|
|
number of elements |
length occupied |
percentage of sequence |
|
|
number of elements |
length occupied |
percentage of sequence |
||
|
SINEs: |
79 |
4,590 bp |
0.00% |
|
SINEs: |
3 |
178 bp |
0.00% |
||
|
|
ALUs |
0 |
0 bp |
0.00% |
|
|
ALUs |
0 |
0 bp |
0.00% |
|
|
MIRs |
65 |
3,731 bp |
0.00% |
|
|
MIRs |
2 |
124 bp |
0.00% |
|
LINEs: |
2,195 |
358,387 bp |
0.19% |
|
LINEs: |
43 |
4,931 bp |
0.03% |
||
|
|
LINE1 |
54 |
3,300 bp |
0.00% |
|
|
LINE1 |
2 |
135 bp |
0.00% |
|
|
LINE2 |
26 |
1,508 bp |
0.00% |
|
|
LINE2 |
2 |
193 bp |
0.00% |
|
|
L3/CR1 |
1,949 |
330,647 bp |
0.17% |
|
|
L3/CR1 |
35 |
4,387 bp |
0.03% |
|
LTR elements: |
303 |
22,297 bp |
0.01% |
|
LTR elements: |
8 |
936 bp |
0.01% |
||
|
|
ERVL |
19 |
1,246 bp |
0.00% |
|
|
ERVL |
3 |
150 bp |
0.00% |
|
|
ERVL-MaLRs |
5 |
305 bp |
0.00% |
|
|
ERVL-MaLRs |
1 |
61 bp |
0.00% |
|
|
ERV_classI |
229 |
16,722 bp |
0.01% |
|
|
ERV_classI |
1 |
87 bp |
0.00% |
|
|
ERV_classII |
33 |
2,034 bp |
0.00% |
|
|
ERV_classII |
1 |
71 bp |
0.00% |
|
DNA elements: |
1,273 |
151,813 bp |
0.08% |
|
DNA elements: |
58 |
14,499 bp |
0.09% |
||
|
|
hAT-Charlie |
305 |
41,456 bp |
0.02% |
|
|
hAT-Charlie |
8 |
3,595 bp |
0.02% |
|
|
TcMar-Tigger |
743 |
88,342 bp |
0.05% |
|
|
TcMar-Tigger |
43 |
9,434 bp |
0.06% |
|
Unclassified: |
3 |
164 bp |
0.00% |
|
Unclassified: |
0 |
0 bp |
0.00% |
||
|
Total interspersed repeats: |
|
537,251 bp |
0.28% |
|
Total interspersed repeats: |
|
20,544 bp |
0.12% |
||
|
Small RNA: |
414 |
27,650 bp |
0.01% |
|
Small RNA: |
16 |
1,010 bp |
0.01% |
||
|
Satellites: |
1 |
46 bp |
0.00% |
|
Satellites: |
0 |
0 bp |
0.00% |
||
|
Simple repeats: |
0 |
0 bp |
0.00% |
|
Simple repeats: |
0 |
0 bp |
0.00% |
||
|
Low complexity: |
0 |
0 bp |
0.00% |
|
Low complexity: |
0 |
0 bp |
0.00% |
||
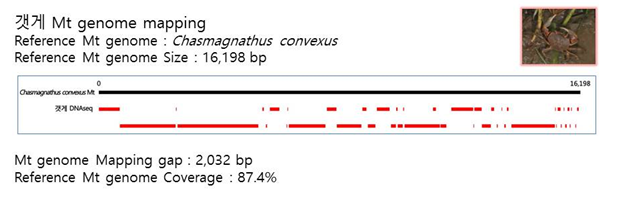
D. Mito-genome Map
(4) Microsatellite 후보군
A. SSR statistics
|
Motif(-mer) |
2 |
3 |
4 |
5 |
6 |
7 |
8 |
9 |
10 |
Total |
|
겟게 |
89,764 |
45,550 |
7,655 |
846 |
60 |
1 |
- |
2 |
1 |
143,879 |
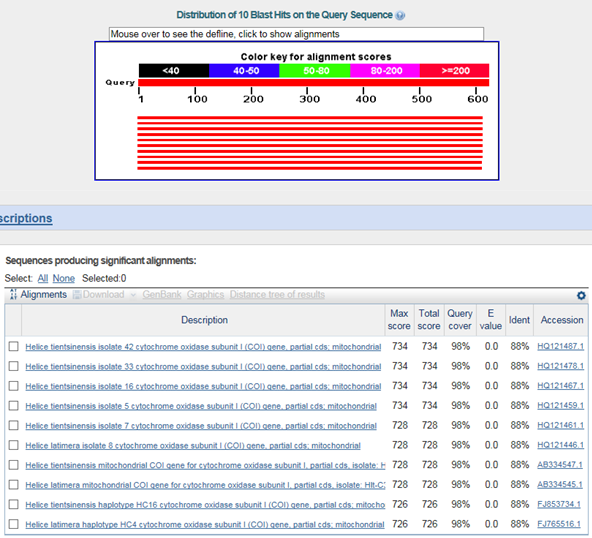
B. 바코드서열
>갯게(Chasmagnathus convexus)
TCGAAAATGGATGTATTTAAATTACGGTCTGTAAGTAGTATAGTAATAGCTCCAGCTAGA
ACTGGCAAGGATAAAAGAAGTAGGATGGCAGTAATAAATACAGCTCAAACAAAAAGGGGT
ATTTGGTCTATCATTATCCCATATGATCGTATATTGATAACGGTTGTTATGAAATTTACA
GCTCCTAGAATTGAGGATACTCCTGCAAGATGAGGAGAAAAAATTCCAAGATCTACGGAG
GCCCCAGCATGGGTGATAGCAGCTGCAAGGGGTGGATAAACAGTTCATCCTGTGCCCACT
CCTCTTTCTACTATTCTTCTTGTTAGGAGAGAAAGAGATGGAGGAAGAAGTCAAAATCTT
ATATTATTTATACGAGGAAAGGCTTTATCTGGTGCTCCTAGTATAAGGGGAACTAGTCAG
TTACCGAATCCTCCAGTTATAATGGGTATAACTATAAAGAAAATTATGACAAAAGCGTGG
GCTGTCACAACCACATTGTAAATTTGGTCATTACCGATTAATCTTCCTGGTTGACTTAGT
TCTGCTCGAATAATTAATCTTACGGATGTACCTACTATTCCGGCTCCAGCGCCGAAAATA
AAATATAAGGTTCCAATGTCTT
갈게(Helice tientsinensis) : family 까지 같음
Helice latimera : family 까지 같음