СІСжЙшВХХаДоЦиРЬ (Aegista (Plectotropis) quelpartensis)
|
|
ЂУ БЙ Иэ: СІСжЙшВХХаДоЦиРЬ ЂУ Ча Иэ: Aegista (Plectotropis) quelpartensis Pilsbry & Hirase ЂУ СіСЄЙјШЃ: АэРЏСО ЂУ Аш Хы: КЙСЗА-СјРЏЦѓИё-ДоЦиРЬАњ |
Ён Л§БшЛѕ : ЦаАЂРК ХОРЬ ГєСі ОЪРК ГЊМБЛѓ ПјЛдИ№ОчРЛ РЬЗщДй. ВЎСњРК КёБГРћ ОуАэ ЙрРК АЅЛіРЛ ЖьИч ЙЋЕђ БЄХУРЬ ГДй. ЦаАЂ РќИщПЁДТ МКРхМБРЛ ЕћЖѓ АЁДТ АИ№АЁ УЮУЮЧЯАд ГЊХИГЊИч УМУў СжПЌПЁДТ ДѕПэ БцОюСЎМ КёДУ И№ОчРЧ Бф АИ№АЁ УМУў АЁРхРкИЎИІ ЕЮИЃАэ РжАэ УМУў ЙиИщПЁЕЕ АЁДТ АИ№АЁ МКРхМБРЛ ЕћЖѓ ПСіОю ГЊХИГДй. ГЊУўРК 7.5Уў РЬИч КРЧеРК БэСі ОЪРИГЊ ЖбЗЧЧЯДй. АЂ ГЊУўРК СЁСјРћРИЗЮ СѕАЁЧЯИч ИЖСіИЗ ГЊУўРЧ АЂБИДТ ОЦЗЁЗЮ УГСјДй. УМУў ОюБњДТ ЧќМКЕЧСі ОЪАэ АЁРхРкИЎРЧ АЂРК АЧЯИч АЂРЛ ЕћЖѓ КёДУИ№ОчРЧ Бф АИ№АЁ РжДй. УМУў ЙиИщРК ЦэЦђЧЯСі ОЪАэ ЕеБлДй. УМУў ГєРЬДТ 8.3mmЗЮ ЦаАЂ ГєРЬРЧ 78%СЄЕЕ РЬДй. АЂБИДТ ХЉАэ УЪНТДо И№ОчРЬИч НЩЧЯАд КёНКЕыЧЯДй. АЂБИ АЁРхРкИЎДТ ЙщЛіРЬДй. ЦаАЂРЧ ХЉБтДТ ГєРЬ 12mm, СіИЇ 21mmРЬДй.
Ён МНФСі : РЮАЁ БйУГ ЕЙДуРЬГЊ ОпЛъПЁ МНФЧбДй.
Ён КаЦї : ЧбБЙ АэРЏСОРИЗЮ СІСжЕЕАЁ И№НФЛъСіРЬИч СІСжЕЕ, АцГВПЁ КаЦїЧбДй.
1) ӪѧѪКИ
- УЄС§ЧуАЁ: АэРЏСО
- УЄ С§ Рк: АПјДыЧаБГ РЬСиЛѓ
2) УЄС§ЧіШВ Йз ЛљЧУУГИЎ
|
СІСжЙшВХХаДоЦиРЬ УЄС§ЧіШВ |
ЛљЧУУГИЎ |
|||
|
УЄС§РЯ |
АГУМ Мі |
DNAseq |
RNAseq |
ЧЅКЛ |
|
2014.08.26 |
14 EA |
13 EA (Йп) |
13 EA (ГЛРх) |
1 EA |
3) QC Data
|
ЛљЧУСО |
DNA QC |
RNA QC |
Result |
||
|
Concentration (ug/ul) |
Total Conc. (ug) |
Concentration (ug/ul) |
RIN (>7) |
||
|
СІСжЙшВХХаДоЦиРЬ |
162 |
17.8 |
393 |
9.5 |
pass |
4) БтСИ NCBI Taxonomy РЏРќСЄКИ ЧіШВ
|
РЏРќСЄКИ СОЗљ |
Aegista Мг РЏРќРкПј |
Aegista (Plectotropis) quelpartensis РЏРќРкПј |
|
Nucleotide |
359 |
- |
|
Protein |
251 |
- |
|
Genome |
2 |
- |
|
Gene |
74 |
- |
5) Sequencing КаМЎ АсАњ
- Illumina DNA Sequncing АсАњ
|
Input information |
|||||
|
ЁЁ |
Region1 |
Region2 |
Region3 |
Region4 |
Total |
|
raw data |
|||||
|
Number Of Reads |
101,887,398 |
101,887,398 |
17,581,833 |
17,581,833 |
238,938,462 |
|
Number Of Bases |
12,837,812,148 |
12,837,812,148 |
2,215,310,958 |
2,215,310,958 |
30,106,246,212 |
|
error correction |
|||||
|
Number Of Reads |
97,446,876 |
95,671,796 |
8,807,996 |
8,464,392 |
210,391,060 |
|
Number Of Bases |
11,085,272,433 |
10,725,990,135 |
870,953,258 |
825,183,376 |
23,507,399,202 |
|
pair reads |
|||||
|
Number Of Reads |
|
184,381,992 |
|
12,007,658 |
196,389,650 |
|
Number Of Bases |
|
20,914,197,839 |
|
1,240,872,266 |
22,155,070,105 |
|
single reads |
|||||
|
Number Of Reads |
|
8,736,680 |
|
5,264,730 |
14,001,410 |
|
Number Of Bases |
|
897,064,729 |
|
455,264,368 |
1,352,329,097 |
|
Assembly Results |
|||||
|
Scaffold Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Scaffolds |
2,277,010 |
2,277,010 |
519,523 |
159,152 |
19,123 |
|
Number Of Bases |
859,264,857 |
859,264,857 |
484,643,590 |
233,650,081 |
47,958,340 |
|
Avg. Scaffold Size |
377 |
377 |
932 |
1,468 |
2,507 |
|
N50 Scaffold Size |
587 |
587 |
976 |
1,443 |
2,421 |
|
N80 Scaffold Size |
231 |
231 |
659 |
1,143 |
2,135 |
|
N90 Scaffold Size |
152 |
152 |
576 |
1,068 |
2,063 |
|
largest Scaffold Size |
9,274 |
9,274 |
9,274 |
9,274 |
9,274 |
|
Contig Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Contigs |
21,566,578 |
3,882,699 |
245,566 |
25,337 |
621 |
|
Number Of Bases |
1,924,868,632 |
927,402,447 |
176,337,472 |
31,720,840 |
1,463,686 |
|
Avg. Contig Size |
89 |
238 |
718 |
1,251 |
2,356 |
|
N50 Contig Size |
91 |
261 |
703 |
1,207 |
2,262 |
|
N80 Contig Size |
49 |
160 |
564 |
1,065 |
2,094 |
|
N90 Contig Size |
47 |
137 |
529 |
1,030 |
2,046 |
|
Largest Contig Size |
8,119 |
8,119 |
8,119 |
8,119 |
8,119 |
- Illumina RNA Sequncing АсАњ
|
Input information |
|||||
|
ЁЁ |
Region1 |
Region2 |
Region3 |
Region4 |
Total |
|
raw data |
|||||
|
Number Of Reads |
101,851,586 |
101,851,586 |
17,769,443 |
17,769,443 |
239,242,058 |
|
Number Of Bases |
12,833,299,836 |
12,833,299,836 |
2,238,949,818 |
2,238,949,818 |
30,144,499,308 |
|
error correction |
|||||
|
Number Of Reads |
100,576,427 |
100,081,232 |
16,991,601 |
16,829,182 |
234,478,442 |
|
Number Of Bases |
12,440,237,553 |
12,420,785,646 |
2,069,164,472 |
2,044,321,924 |
28,974,509,595 |
|
pair reads |
|||||
|
Number Of Reads |
|
199,237,908 |
|
33,153,936 |
232,391,844 |
|
Number Of Bases |
|
24,713,305,393 |
|
4,049,130,552 |
28,762,435,945 |
|
single reads |
|||||
|
Number Of Reads |
|
1,419,751 |
|
666,847 |
2,086,598 |
|
Number Of Bases |
|
147,717,806 |
|
64,355,844 |
212,073,650 |
|
Assembly Results |
|||||
|
Scaffold Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Scaffolds |
319,952 |
319,952 |
39,978 |
12,054 |
2,597 |
|
Number Of Bases |
93,918,909 |
93,918,909 |
39,476,731 |
20,394,949 |
7,723,326 |
|
Avg. Scaffold Size |
293 |
293 |
987 |
1,691 |
2,973 |
|
N50 Scaffold Size |
392 |
392 |
1,028 |
1,676 |
2,903 |
|
N80 Scaffold Size |
164 |
164 |
652 |
1,199 |
2,267 |
|
N90 Scaffold Size |
131 |
131 |
570 |
1,094 |
2,125 |
|
largest Scaffold Size |
17,321 |
17,321 |
17,321 |
17,321 |
17,321 |
|
Contig Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Contigs |
2,954,097 |
437,327 |
27,246 |
4,975 |
614 |
|
Number Of Bases |
254,535,026 |
98,293,159 |
22,112,723 |
7,339,704 |
1,651,717 |
|
Avg. Contig Size |
86 |
224 |
811 |
1,475 |
2,690 |
|
N50 Contig Size |
91 |
242 |
790 |
1,421 |
2,580 |
|
N80 Contig Size |
52 |
143 |
586 |
1,130 |
2,162 |
|
N90 Contig Size |
47 |
120 |
540 |
1,059 |
2,095 |
|
Largest Contig Size |
8,859 |
8,859 |
8,859 |
8,859 |
8,859 |
ГЊ) ИъСОРЇБтСО ЕПЙА Genome Survey
|
АЫМіПы ЛљЧУ DNA ГѓЕЕ Йз Уб Оч, ШЎСѕЧЅКЛ ЧіШВЧЅ |
|||||||
|
ЁЁSampleИэ |
ГѓЕЕ(ng/ul) |
Volume(ul) |
DNA УбЗЎ(ug) |
O.D A260/280 |
ШЎСѕЧЅКЛ |
УЄС§ЧуАЁРЯ |
УЄС§РЯ |
|
СІСжЙшВХХаДоЦиРЬ |
104 |
996 |
103 |
1.72 |
Ёл |
АэРЏСО |
2014.8.26 |
(2) РЏРќУМ МПШЎКИ : ЧіРч 20СО ШЎКИ
|
Sample |
БИКа |
raw data size |
trim data size |
КёАэ |
|
СІСжЙшВХХаДоЦиРЬ |
DNA |
30,106,246,212 |
23,507,399,202 |
ЁЁ |
|
RNA |
30,144,499,308 |
28,974,509,595 |
ЁЁ |
(3) БтХИКаМЎ
A. Genome Size estimation
|
Лљ ЧУ Иэ |
Kmer size |
Kmer total |
Pk depth |
Genome_size |
Coverage(X) |
|
СІСжЙшВХХаДоЦиРЬ |
23 |
15,894,434,088 |
147 |
108,125,401 |
188.462 |
B. ЙнКЙМП
- НУФіНЬРЬ ПЯЗсЕШ СІСжЙшВХХаДоЦиРЬ DNA Йз RNAРЧ contigs Сп 1kb РЬЛѓРЧ contigsРЧ ЙнКЙМПРЛ ХНЛі(Left; Repeat masking results of DNA contigs, Right; Repeat masking results of RNA contigs).
|
sequences: 26,032 |
|
sequences: 7,108 |
||||||||
|
total length: 32,608,921 bp (32,608,921 bp excl N/X-runs) |
|
total length: 11,220,308 bp (11,220,308 bp excl N/X-runs) |
||||||||
|
GC level: 40.72% |
|
GC level: 44.87% |
||||||||
|
bases masked: 54749 bp (0.17%) |
|
bases masked: 7823 bp (0.07%) |
||||||||
|
|
number of elements |
length occupied |
percentage of sequence |
|
|
number of elements |
length occupied |
percentage of sequence |
||
|
SINEs: |
5 |
229 bp |
0.00% |
|
SINEs: |
0 |
0 bp |
0.00% |
||
|
|
ALUs |
0 |
0 bp |
0.00% |
|
|
ALUs |
0 |
0 bp |
0.00% |
|
|
MIRs |
2 |
95 bp |
0.00% |
|
|
MIRs |
0 |
0 bp |
0.00% |
|
LINEs: |
105 |
25,847 bp |
0.08% |
|
LINEs: |
12 |
1,931 bp |
0.02% |
||
|
|
LINE1 |
7 |
395 bp |
0.00% |
|
|
LINE1 |
2 |
87 bp |
0.00% |
|
|
LINE2 |
3 |
134 bp |
0.00% |
|
|
LINE2 |
1 |
115 bp |
0.00% |
|
|
L3/CR1 |
14 |
879 bp |
0.00% |
|
|
L3/CR1 |
3 |
160 bp |
0.00% |
|
LTR elements: |
17 |
2,171 bp |
0.01% |
|
LTR elements: |
4 |
577 bp |
0.01% |
||
|
|
ERVL |
0 |
0 bp |
0.00% |
|
|
ERVL |
0 |
0 bp |
0.00% |
|
|
ERVL-MaLRs |
0 |
0 bp |
0.00% |
|
|
ERVL-MaLRs |
1 |
66 bp |
0.00% |
|
|
ERV_classI |
13 |
1,802 bp |
0.00% |
|
|
ERV_classI |
2 |
203 bp |
0.00% |
|
|
ERV_classII |
1 |
39 bp |
0.00% |
|
|
ERV_classII |
0 |
0 bp |
0.00% |
|
DNA elements: |
179 |
20,437 bp |
0.06% |
|
DNA elements: |
28 |
4,899 bp |
0.04% |
||
|
|
hAT-Charlie |
61 |
7,320 bp |
0.02% |
|
|
hAT-Charlie |
15 |
3,017 bp |
0.03% |
|
|
TcMar-Tigger |
29 |
3,092 bp |
0.01% |
|
|
TcMar-Tigger |
2 |
105 bp |
0.00% |
|
Unclassified: |
2 |
102 bp |
0.00% |
|
Unclassified: |
0 |
0 bp |
0.00% |
||
|
Total interspersed repeats: |
|
48,786 bp |
0.15% |
|
Total interspersed repeats: |
|
7,407 bp |
0.07% |
||
|
Small RNA: |
52 |
5,963 bp |
0.02% |
|
Small RNA: |
2 |
416 bp |
0.00% |
||
|
Satellites: |
0 |
0 bp |
0.00% |
|
Satellites: |
0 |
0 bp |
0.00% |
||
|
Simple repeats: |
0 |
0 bp |
0.00% |
|
Simple repeats: |
0 |
0 bp |
0.00% |
||
|
Low complexity: |
0 |
0 bp |
0.00% |
|
Low complexity: |
0 |
0 bp |
0.00% |
||
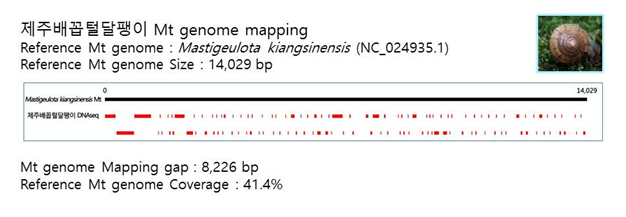
D. Mito-genome Map
(4) Microsatellite ШФКИБК
A. SSR statistics
|
Motif(-mer) |
2 |
3 |
4 |
5 |
6 |
7 |
8 |
9 |
10 |
Total |
|
СІСжЙшВХХаДоЦиРЬ |
4,423 |
1,950 |
566 |
11 |
3 |
1 |
2 |
1 |
- |
6,957 |
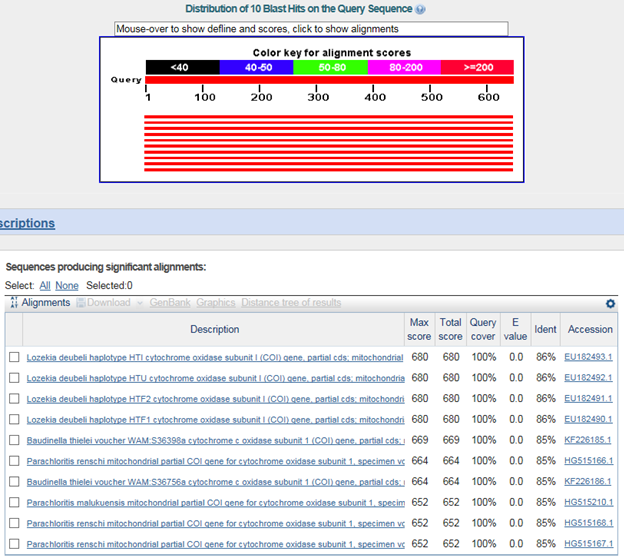
B. ЙйФкЕхМП
>СІСжЙшВХХаДоЦиРЬ(Aegista quelpartensis)
TTATATATATTATTTGGTGTATGATGTGGTATAGTAGGATCTGGTTTATCAATATTAATTCGTATAGAACTAGGGACTTGTAATGTATTAAGGGATAGTCATTTTTTTAATGTAATCGTTACAGCTCATGCATTTGTAATAATTTTTTTTATAGTAATACCTATTATAATTGGAGGTTTTGGAAATTGAATAGTTCCTTTATTAATTGGAGCTCCTGATATAAGTTTTCCACGAATAAATAATATAAGCTTTTGATTATTACCTCCTTCATTTATTTTATTAATTTCTAGAAGGTTAGTTGAGGGTGGAGCTGGTACTGGATGAACAGTATATCCACCTTTAAGTTCTATTGTAGGTCATAGAGGAGCCTCAGTAGATTTAGCCATTTTTTCTCTGCATTTGGCTGGTATATCTTCAATTCTAGGTGCTATTAATTTTATTACAACAATTTTTAATATGCGTAGTCCAGGAATTTCATTAGAACGAATAAGTTTATTTGTATGATCTATTTTAATTACTGTATTTTTATTACTTTTATCTTTACCTGTTTTAGCAGGTGCTATTACTATATTGCTTACTGATCGAAATTTTAATACTTCTTTCTTTGATCCTGCAGGCGGAGGAGATCCAATTTTATATCAGCATTT
Lozekia deubeli : family КЮХЭ ДйИЇ
Baudinella thielei :ЁЁsuperfamily КЮХЭ ДйИЇ