제주배꼽달팽이 (Aegista (Aegista) chejuensis)
|
|
▣ 국 명: 제주배꼽달팽이 ▣ 학 명: Aegista chejuensis Pilsbry & Hirase ▣ 지정번호: 고유종 ▣ 계 통: 복족강-진유폐목-달팽이과 |
◎ 생김새 : 패각은 나탑이 낮지 않은 나선상의 원뿔모양이며 껍질은 비교적 단단하다. 패각 전면에는 광택이 나는 연한 갈색의 각피로 덮여 있고, 매우 가는 실모양의 굴곡이 성장선을 따라 나선상으로 조밀하게 뻗어 있다. 밑면은 가늘고 촘촘한 성장선이 나타나고 있으며 굴곡이 심하지 않아서 패각 전면보다 매끄럽다. 크기는 높이 6.2mm, 지름 10.2mm다. 나층은 6~6.5층이며 황갈색을 띤다. 각 나층의 크기는 규칙적으로 조금씩 증가한다. 마지막 나층의 각구는 아래로 처지지 않으며 체층 가장자리는 각이 없어 둥글다. 각구는 패각에 비하여 비교적 크고 비스듬하며 둥근 초승달모양이다.
◎ 서식지 : 숲 속의 다소 건조한 관목림 밑이나 돌무덤 속에 서식한다.
◎ 분포 : 한국 고유종으로 모식산지는 제주도이며 진도, 남해안에 분포한다.
1) 채집정보
- 채집허가: 고유종
- 채 집 자: 강원대학교 이준상
2) 채집현황 및 샘플처리
|
제주배꼽달팽이 채집현황 |
샘플처리 |
|||
|
채집일 |
개체 수 |
DNAseq |
RNAseq |
표본 |
|
2014.08.26 |
9 EA |
4 EA |
4 EA |
1 EA |
3) QC Data
|
샘플종 |
DNA QC |
RNA QC |
Result |
||
|
Concentration (ug/ul) |
Total Conc. (ug) |
Concentration (ug/ul) |
RIN (>7) |
||
|
제주배꼽달팽이 |
113 |
9 |
254 |
9.8 |
pass |
4) 기존 NCBI Taxonomy 유전정보 현황
|
유전정보 종류 |
Aegista 속 유전자원 |
Aegista (Aegista) chejuensis 유전자원 |
|
Nucleotide |
359 |
- |
|
Protein |
251 |
- |
|
Genome |
2 |
- |
|
Gene |
74 |
- |
5) Sequencing 분석 결과
- Illumina DNA Sequncing 결과
|
Input information |
|||||
|
|
Region1 |
Region2 |
Region3 |
Region4 |
Total |
|
raw data |
|||||
|
Number Of Reads |
118,506,288 |
118,506,288 |
10,275,102 |
10,275,102 |
257,562,780 |
|
Number Of Bases |
14,931,792,288 |
14,931,792,288 |
1,294,662,852 |
1,294,662,852 |
32,452,910,280 |
|
error correction |
|||||
|
Number Of Reads |
109,658,743 |
108,690,436 |
4,491,029 |
4,422,673 |
227,262,881 |
|
Number Of Bases |
12,563,157,322 |
12,348,029,788 |
456,363,875 |
444,403,694 |
25,811,954,679 |
|
pair reads |
|||||
|
Number Of Reads |
|
209,527,066 |
|
6,725,334 |
216,252,400 |
|
Number Of Bases |
|
24,031,628,914 |
|
713,837,970 |
24,745,466,884 |
|
single reads |
|||||
|
Number Of Reads |
|
8,822,113 |
|
2,188,368 |
11,010,481 |
|
Number Of Bases |
|
879,558,196 |
|
186,929,599 |
1,066,487,795 |
|
Assembly Results |
|||||
|
Scaffold Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Scaffolds |
2,215,276 |
2,215,276 |
511,727 |
186,167 |
37,418 |
|
Number Of Bases |
886,619,538 |
886,619,538 |
540,436,120 |
313,208,676 |
111,951,531 |
|
Avg. Scaffold Size |
400 |
400 |
1,056 |
1,682 |
2,991 |
|
N50 Scaffold Size |
680 |
680 |
1,137 |
1,659 |
2,791 |
|
N80 Scaffold Size |
240 |
240 |
703 |
1,208 |
2,238 |
|
N90 Scaffold Size |
148 |
148 |
595 |
1,097 |
2,111 |
|
largest Scaffold Size |
99,405 |
99,405 |
99,405 |
99,405 |
99,405 |
|
Contig Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Contigs |
21,540,436 |
4,140,248 |
258,618 |
29,414 |
2,596 |
|
Number Of Bases |
1,961,131,938 |
968,202,411 |
195,406,419 |
45,440,146 |
12,182,373 |
|
Avg. Contig Size |
91 |
233 |
755 |
1,544 |
4,692 |
|
N50 Contig Size |
96 |
249 |
717 |
1,347 |
5,704 |
|
N80 Contig Size |
50 |
155 |
568 |
1,096 |
2,685 |
|
N90 Contig Size |
47 |
134 |
532 |
1,045 |
2,240 |
|
Largest Contig Size |
70,769 |
70,769 |
70,769 |
70,769 |
70,769 |
- Illumina RNA Sequncing 결과
|
Input information |
|||||
|
|
|
|
Region1 |
Region2 |
Total |
|
raw data |
|||||
|
Number Of Reads |
|
|
128,327,935 |
128,327,935 |
256,655,870 |
|
Number Of Bases |
|
|
16,169,319,810 |
16,169,319,810 |
32,338,639,620 |
|
error correction |
|||||
|
Number Of Reads |
|
|
127,109,211 |
126,147,475 |
253,256,686 |
|
Number Of Bases |
|
|
15,739,923,181 |
15,660,526,491 |
31,400,449,672 |
|
pair reads |
|||||
|
Number Of Reads |
|
|
|
|
251,314,118 |
|
Number Of Bases |
|
|
|
|
31,188,107,081 |
|
single reads |
|||||
|
Number Of Reads |
|
|
|
|
1,942,568 |
|
Number Of Bases |
|
|
|
|
212,342,591 |
|
Assembly Results |
|||||
|
Scaffold Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Scaffolds |
228,106 |
228,106 |
34,622 |
11,146 |
2,646 |
|
Number Of Bases |
74,648,423 |
74,648,423 |
35,620,275 |
19,569,119 |
8,016,049 |
|
Avg. Scaffold Size |
327 |
327 |
1,028 |
1,755 |
3,029 |
|
N50 Scaffold Size |
467 |
467 |
1,096 |
1,763 |
2,940 |
|
N80 Scaffold Size |
189 |
189 |
667 |
1,233 |
2,278 |
|
N90 Scaffold Size |
134 |
134 |
576 |
1,108 |
2,128 |
|
largest Scaffold Size |
14,943 |
14,943 |
14,943 |
14,943 |
14,943 |
|
Contig Metrics |
> 1 bp* |
> 100 bp |
> 500 bp** |
> 1000 bp |
> 2000 bp |
|
Number Of Contigs |
3,795,144 |
321,887 |
22,581 |
5,292 |
722 |
|
Number Of Bases |
230,294,252 |
74,003,080 |
19,581,172 |
7,973,399 |
1,940,159 |
|
Avg. Contig Size |
60 |
229 |
867 |
1,506 |
2,687 |
|
N50 Contig Size |
63 |
261 |
870 |
1,472 |
2,638 |
|
N80 Contig Size |
35 |
138 |
607 |
1,148 |
2,164 |
|
N90 Contig Size |
32 |
115 |
549 |
1,066 |
2,077 |
|
Largest Contig Size |
7,490 |
7,490 |
7,490 |
7,490 |
7,490 |
나) 멸종위기종 동물 Genome Survey
|
검수용 샘플 DNA 농도 및 총 양, 확증표본 현황표 |
|||||||
|
Sample명 |
농도(ng/ul) |
Volume(ul) |
DNA 총량(ug) |
O.D A260/280 |
확증표본 |
채집허가일 |
채집일 |
|
제주배꼽달팽이 |
152 |
996 |
151 |
1.54 |
○ |
고유종 |
2014.8.26 |
(2) 유전체 서열확보 : 현재 20종 확보
|
Sample |
구분 |
raw data size |
trim data size |
비고 |
|
제주배꼽달팽이 |
DNA |
32,452,910,280 |
25,811,954,679 |
|
|
RNA |
32,338,639,620 |
31,400,449,672 |
|
(3) 기타분석
A. Genome Size estimation
|
샘 플 명 |
Kmer size |
Kmer total |
Pk depth |
Genome_size |
Coverage(X) |
|
제주배꼽달팽이 |
23 |
18,486,980,928 |
178 |
103,859,443 |
228.205 |
B. 반복서열
- 시퀀싱이 완료된 제주배꼽달팽이 DNA 및 RNA의 contigs 중 1kb 이상의 contigs의 반복서열을 탐색(Left; Repeat masking results of DNA contigs, Right; Repeat masking results of RNA contigs).
|
sequences: 29,640 |
|
sequences: 5,292 |
||||||||
|
total length: 45736033 bp (45736033 bp excl N/X-runs) |
|
total length: 7,973,399 bp (7,973,399 bp excl N/X-runs) |
||||||||
|
GC level: 41.78% |
|
GC level: 45.61% |
||||||||
|
bases masked: 60,083 bp (0.13%) |
|
bases masked: 4079 bp (0.05%) |
||||||||
|
|
number of elements |
length occupied |
percentage of sequence |
|
|
number of elements |
length occupied |
percentage of sequence |
||
|
SINEs: |
8 |
367 bp |
0.00% |
|
SINEs: |
0 |
0 bp |
0.00% |
||
|
|
ALUs |
0 |
0 bp |
0.00% |
|
|
ALUs |
0 |
0 bp |
0.00% |
|
|
MIRs |
3 |
178 bp |
0.00% |
|
|
MIRs |
0 |
0 bp |
0.00% |
|
LINEs: |
106 |
24,847 bp |
0.05% |
|
LINEs: |
14 |
1,154 bp |
0.01% |
||
|
|
LINE1 |
9 |
584 bp |
0.00% |
|
|
LINE1 |
4 |
245 bp |
0.00% |
|
|
LINE2 |
4 |
223 bp |
0.00% |
|
|
LINE2 |
3 |
146 bp |
0.00% |
|
|
L3/CR1 |
14 |
1,645 bp |
0.00% |
|
|
L3/CR1 |
1 |
79 bp |
0.00% |
|
LTR elements: |
24 |
2,414 bp |
0.01% |
|
LTR elements: |
1 |
38 bp |
0.00% |
||
|
|
ERVL |
1 |
63 bp |
0.00% |
|
|
ERVL |
0 |
0 bp |
0.00% |
|
|
ERVL-MaLRs |
0 |
0 bp |
0.00% |
|
|
ERVL-MaLRs |
0 |
0 bp |
0.00% |
|
|
ERV_classI |
18 |
1,872 bp |
0.00% |
|
|
ERV_classI |
1 |
38 bp |
0.00% |
|
|
ERV_classII |
0 |
0 bp |
0.00% |
|
|
ERV_classII |
0 |
0 bp |
0.00% |
|
DNA elements: |
177 |
20,735 bp |
0.05% |
|
DNA elements: |
18 |
2,687 bp |
0.03% |
||
|
|
hAT-Charlie |
60 |
7,524 bp |
0.02% |
|
|
hAT-Charlie |
10 |
2,150 bp |
0.03% |
|
|
TcMar-Tigger |
42 |
5,299 bp |
0.01% |
|
|
TcMar-Tigger |
2 |
71 bp |
0.00% |
|
Unclassified: |
2 |
94 bp |
0.00% |
|
Unclassified: |
0 |
0 bp |
0.00% |
||
|
Total interspersed repeats: |
|
48,457 bp |
0.11% |
|
Total interspersed repeats: |
|
3,879 bp |
0.05% |
||
|
Small RNA: |
99 |
11,300 bp |
0.02% |
|
Small RNA: |
2 |
200 bp |
0.00% |
||
|
Satellites: |
2 |
326 bp |
0.00% |
|
Satellites: |
0 |
0 bp |
0.00% |
||
|
Simple repeats: |
0 |
0 bp |
0.00% |
|
Simple repeats: |
0 |
0 bp |
0.00% |
||
|
Low complexity: |
0 |
0 bp |
0.00% |
|
Low complexity: |
0 |
0 bp |
0.00% |
||
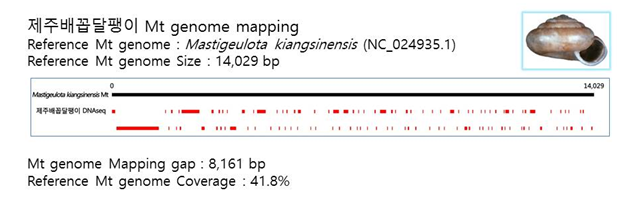
D. Mito-genome Map
(4) Microsatellite 후보군
A. SSR statistics
|
Motif(-mer) |
2 |
3 |
4 |
5 |
6 |
7 |
8 |
9 |
10 |
Total |
|
제주배꼽달팽이 |
5,633 |
2,311 |
626 |
17 |
14 |
5 |
7 |
1 |
3 |
8,617 |
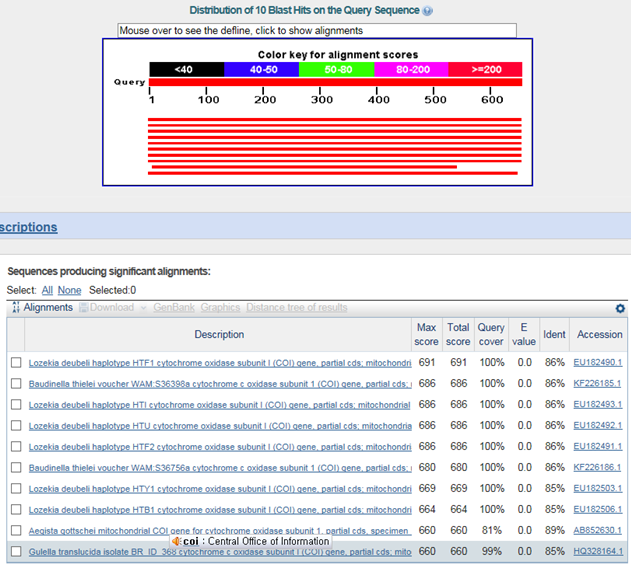
B. 바코드서열
>제주배꼽달팽이(Aegista chejuensis)
ACTTTATATATATTATTTGGTGTTTGATGTGGAATAGTTGGATCTGGTTTATCATTATTA
ATCCGTATAGAATTAGGAACTTGTAATGTATTAAGCGATAGTCATTTTTTTAATGTTATT
GTTACAGCACATGCATTTGTAATAATCTTCTTTATAGTAATACCAATCATGATTGGTGGT
TTTGGAAACTGAATAGTTCCATTACTAATTGGGGCTCCTGATATAAGTTTTCCACGAATA
AATAATATAAGATTTTGATTATTACCACCTTCATTTATCTTATTAATTTCAAGAAGATTA
GTTGAAGGTGGGGCTGGAACTGGGTGAACTGTATATCCACCTTTAAGTTCAATTATAGGT
CATAGAGGTGCTTCAGTAGATTTAGCAATTTTTTCTTTACATTTAGCTGGAATATCATCT
ATTTTAGGTGCAATTAATTTTATTACTACAATTTTTAATATACGTAGGCCAGGAAACTCT
ATAGAACGATTAAGATTATTTGTATGATCTATTTTAATTACAGTATTTTTATTACTTCTA
TCATTACCTGTTTTAGCAGGTGCTATTACAATATTATTAACTGATCGAAATTTTAATACT
TCATTTTTTGATCCTGCAGGAGGAGGAGATCCTATTTTATATCAGCATTT
Lozekia deubeli : family 부터 다름
Baudinella thielei : superfamily 부터 다름